| Line 821: | Line 821: | ||
<h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Characterisation </h2> | <h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Characterisation </h2> | ||
| − | <p>The expression of deGFP was | + | <p>The expression of deGFP was first tested in E. coli cells using an experimental procedure similar to that used in the Interlab study. Cells transformed with pSB1C3-J23100-deGFP were grown in 10 mL LB broth overnight and OD600 nm was measured. Culture was added to 3 separate falcon tubes and made up to 12 mL with LB with chloramphenicol such that the starting OD600 of the culture was approximately 0.02. This set-up was repeated with cells containing an identical plasmid and construct, except sfGFP was in place of deGFP. As a control, untransformed cells were also prepared identically except the LB did not contain chloramphenicol. Tubes with only LB and LB+chloramphenicol were also prepared as blanks. |
</br></br> | </br></br> | ||
The cultures were shake-incubated at 37oC. 300 μL samples from each tube were taken at time points of 15 mins, 2 hours, 4 hours, and 6 hours and stored at 4oC until the end of the experiment. 100 μL of each sample were then added to a 96-well plate. Fluorescence (excitation 485 nm, emission 510 nm) and absorbance (OD600 nm) were measured using a BMG-Labtech fluostar optima plate reader. Fluorescence intensity and grow rates for all three cell types were calculated over time (Figure 1). It was found that while cells expressing sfGFP had a much higher fluorescence intensity than cells expressing deGFP, the growth rate for cells with deGFP was closer to that of untransformed cells. This suggests that in vivo, either deGFP has lower expression than sfGFP, or each molecule of deGFP emits less fluorescence. | The cultures were shake-incubated at 37oC. 300 μL samples from each tube were taken at time points of 15 mins, 2 hours, 4 hours, and 6 hours and stored at 4oC until the end of the experiment. 100 μL of each sample were then added to a 96-well plate. Fluorescence (excitation 485 nm, emission 510 nm) and absorbance (OD600 nm) were measured using a BMG-Labtech fluostar optima plate reader. Fluorescence intensity and grow rates for all three cell types were calculated over time (Figure 1). It was found that while cells expressing sfGFP had a much higher fluorescence intensity than cells expressing deGFP, the growth rate for cells with deGFP was closer to that of untransformed cells. This suggests that in vivo, either deGFP has lower expression than sfGFP, or each molecule of deGFP emits less fluorescence. | ||
| Line 887: | Line 887: | ||
<div id="chromoproteins" class="collapse"> | <div id="chromoproteins" class="collapse"> | ||
| − | <h2 style="font-size: 1em"> BioBricks used: <a href="http://parts.igem.org/Part:BBa_K2205016">BBa_K2205016(New)</a>,<a href="http://parts.igem.org/Part:BBa_K2205017">BBa_K2205017(New)</a> | + | <h2 style="font-size: 1em"> BioBricks used: <a href="http://parts.igem.org/Part:BBa_K2205016">BBa_K2205016(New)</a>,<a href="http://parts.igem.org/Part:BBa_K2205017">BBa_K2205017(New)</a>,<a href="http://parts.igem.org/Part:BBa_K2205018">BBa_K2205018(New)</a>, <a href="http://parts.igem.org/Part:BBa_K1033915">BBa_K1033915(Uppsala 2013)</a>, <a href="http://parts.igem.org/Part:BBa_K1033925">BBa_K1033925 (Uppsala 2013)</a>, <a href="http://parts.igem.org/Part:BBa_K1033929">BBa_K1033929 (Uppsala 2013)</a> </h2> |
<h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Rationale and Aim </h2> | <h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Rationale and Aim </h2> | ||
| − | <p>The Sensynova multicellular biosensor platform has been developed to overcome the limitations identified by our team | + | <p>The Sensynova multicellular biosensor platform has been developed to overcome the <a href="https://2017.igem.org/Team:Newcastle/HP/Silver">limitations identified by our team</a> that hamper success in biosensor development. One of these limits regards the lack of modularity and reusability of the various components. Our platform design, based on the expression of three main modules (Detector, Processor and Reporter) by three <i>E.coli</i> strains in co-culture, allows the switch of possible variances for each module and the production of multiple customised biosensors. |
</br></br> | </br></br> | ||
This section of the project is based on testing the modularity of the system by replacing the sfGFP output part of the Sensynova platform design with three different output chromoprotein variants; BBa_K1033929 (aeBlue), BBa_K1033925 (spisPink) and BBa_K1033915 (amajLime).</p> | This section of the project is based on testing the modularity of the system by replacing the sfGFP output part of the Sensynova platform design with three different output chromoprotein variants; BBa_K1033929 (aeBlue), BBa_K1033925 (spisPink) and BBa_K1033915 (amajLime).</p> | ||
| Line 963: | Line 963: | ||
<h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Background Information </h2> | <h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Background Information </h2> | ||
<p><b>Human Practices Quotes: </b></p> | <p><b>Human Practices Quotes: </b></p> | ||
| − | <p><b>Biosensor Development.</b> When developing biosensors, it would be useful to test multiple variants of a circuit. This is especially important in the fine-tuning of biosensor behaviour as this requires the screening of many variants to find appropriate activation thresholds for a system. Apart from the initial detection unit, many constructs used in synthetic biology based biosensors are the reusable between different biosensor systems, such as fluorescent protein coding sequences or devices which amplify signals. However, these parts rarely get reused. For example, the Cambridge iGEM | + | <p><b>Biosensor Development.</b> When developing biosensors, it would be useful to test multiple variants of a circuit. This is especially important in the fine-tuning of biosensor behaviour as this requires the screening of many variants to find appropriate activation thresholds for a system. Apart from the initial detection unit, many constructs used in synthetic biology based biosensors are the reusable between different biosensor systems, such as fluorescent protein coding sequences or devices which amplify signals. However, these parts rarely get reused. For example, the Cambridge iGEM team (2009) developed a library of sensitivity tuners which were able to convert polymerase per second inputs to a desired polymerase per second output, allowing a biosensor developer control over the sensitivity of their systems to various target analyte concentrations. This project was impressive enough to win the competition. However, despite the parts' clear usefulness, there is no documentation that the parts have ever been successfully reused within the iGEM competition. We suggest that this is due to the difficulties in assembling biosensors systems – the screening of a library of sensitivity tuners would require the ability to easily generate multiple sensor circuits. Although only one part would be changing in each circuit variant, current genetic engineering techniques mean that parts are tightly coupled together, preventing the simple swapping of parts. |
</br></br> | </br></br> | ||
Therefore, we propose a modular, multicellular system for biosensor development, using a cell-to-cell communication system to eradicate the requirement for further genetic engineering of reusable biosensor devices (Figure 1). | Therefore, we propose a modular, multicellular system for biosensor development, using a cell-to-cell communication system to eradicate the requirement for further genetic engineering of reusable biosensor devices (Figure 1). | ||
| Line 977: | Line 977: | ||
<h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Cell-to-Cell communication </h2> | <h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Cell-to-Cell communication </h2> | ||
| − | <p>Bacteria have native quorum sensing systems which enable cell-to-cell communication through the production and detection of hormone-like auto-inducers. These molecules allow the synchronisation of behaviour in large populations of bacterial cells (Waters & Bassler, 2005). One such system involves the autoinducer AHL (Acylated Homoserine Lactone). AHLs compose of a lactone ring with an acyl side chain containing between 4 and 18 carbons (Churchill & Chen, 2011). Various AHL synthases | + | <p>Bacteria have native quorum sensing systems which enable cell-to-cell communication through the production and detection of hormone-like auto-inducers. These molecules allow the synchronisation of behaviour in large populations of bacterial cells (Waters & Bassler, 2005). One such system involves the autoinducer AHL (Acylated Homoserine Lactone). AHLs compose of a lactone ring with an acyl side chain containing between 4 and 18 carbons (Churchill & Chen, 2011). Various AHL synthases exist, which produce AHL with different modifications and side change lengths. AHL receptors are sensitive to AHLs of specific length. For example, it has been found that the Rhl system, producing and detecting AHL of acyl carbon length 4 and the Las system, producing and detecting AHL of acyl carbon length 12, exhibit little crosstalk – the receptor component of the system is sensitive only to carbon chains of the correct length (Brenner et al., 2007). The orthogonal nature of the AHL family of autoinducers has enabled their use in a variety of synthetic systems. They are often used as biological “wires”, linking either inter- or intracellular processes. These “wires” have been previously used in a number of synthetic biology systems, e.g. Gupta et al. (2013) and Tasmir et al. (2011). |
</br></br> | </br></br> | ||
In this project, it is proposed that modularity, and therefore the ability to use parts “off-the-shelf” without further genetic engineering, could be improved by splitting components of biosensors into different cells which communicate to coordinate responses. The orthogonal quorum sensing systems Rhl and Las will be used as biological “wires”, linking different biosensor components together. This separation of components will enable the decoupling of non-specific components from specific detection systems. Using this approach, production of biosensor variants will not require subsequent engineering steps: cells containing desired components will simply be mixed together (Figure 2). | In this project, it is proposed that modularity, and therefore the ability to use parts “off-the-shelf” without further genetic engineering, could be improved by splitting components of biosensors into different cells which communicate to coordinate responses. The orthogonal quorum sensing systems Rhl and Las will be used as biological “wires”, linking different biosensor components together. This separation of components will enable the decoupling of non-specific components from specific detection systems. Using this approach, production of biosensor variants will not require subsequent engineering steps: cells containing desired components will simply be mixed together (Figure 2). | ||
| Line 1,016: | Line 1,016: | ||
<h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Implementation </h2> | <h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Implementation </h2> | ||
| − | <p>To prove that our concept of splitting biosensors across multiple cells would work, we designed an IPTG sensor. The design of this system can be found in Figure 4. In this system, LacI is constitutively expressed in the detector cell and represses the production of LasI. When IPTG is added, it binds LacI, preventing repression. Therefore, in the presence of IPTG, LasI will produce C12, our first connector molecule. picture To determine that our system would work, it was first tested in silico. Details on the model of this system can be found on our <a href="https://2017.igem.org/Team:Newcastle/ | + | <p>To prove that our concept of splitting biosensors across multiple cells would work, we designed an IPTG sensor. The design of this system can be found in Figure 4. In this system, LacI is constitutively expressed in the detector cell and represses the production of LasI. When IPTG is added, it binds LacI, preventing repression. Therefore, in the presence of IPTG, LasI will produce C12, our first connector molecule. picture To determine that our system would work, it was first tested in silico. Details on the model of this system can be found on our <a href="https://2017.igem.org/Team:Newcastle/Model">Modelling page</a>. |
</br></br> | </br></br> | ||
<img src="https://static.igem.org/mediawiki/2017/5/5c/Iptg_framework.jpg" class="img-fluid rounded mx-auto d-block" style="max-width: 60%" alt=""> | <img src="https://static.igem.org/mediawiki/2017/5/5c/Iptg_framework.jpg" class="img-fluid rounded mx-auto d-block" style="max-width: 60%" alt=""> | ||
| Line 1,023: | Line 1,023: | ||
</br></br> | </br></br> | ||
| − | <p>Parts were synthesised by IDT and integration into the pSB1C3 plasmid confirmed by colony PCR and subsequent sequencing. Red boxes show | + | <p>Parts were synthesised by IDT and integration into the pSB1C3 plasmid confirmed by colony PCR and subsequent sequencing. Red boxes show parts later used for biobrick production (Figure 5). </p> |
<img src="https://static.igem.org/mediawiki/2017/1/10/Framework_gel_parts.png" class="img-fluid rounded mx-auto d-block" style="max-width: 60%" alt=""> | <img src="https://static.igem.org/mediawiki/2017/1/10/Framework_gel_parts.png" class="img-fluid rounded mx-auto d-block" style="max-width: 60%" alt=""> | ||
| Line 1,071: | Line 1,071: | ||
</p> </br> | </p> </br> | ||
<p> | <p> | ||
| − | Results from the modelling predicted that the 1:1:1 ratio is not the optimal combination for the Sensynova device to work. It is | + | Results from the modelling predicted that the 1:1:1 ratio is not the optimal combination for the Sensynova device to work. It is in fact suggested to adopt a higher concentration of the reporter culture compare with the detector and processor. Thus, the framework test was repeated with the 1:1:13 cultures combination. The experiment results, shown in the picture below, confirm the modelling data. There is a consistent discrepancy between IPTG induced and non-induced samples in the 1:1:13 co-cultures, in comparison with the 1:1:1 co-cultures which don't show any difference in presence or absence of IPTG (figure9).</p> |
<p> | <p> | ||
<img src="https://static.igem.org/mediawiki/2017/9/9e/Ratios.jpg" class="img-fluid rounded mx-auto d-block" style="max-width: 60%" alt=""> | <img src="https://static.igem.org/mediawiki/2017/9/9e/Ratios.jpg" class="img-fluid rounded mx-auto d-block" style="max-width: 60%" alt=""> | ||
| Line 1,151: | Line 1,151: | ||
</br></br> | </br></br> | ||
Waters, C. & Bassler, B. (2005) Quorum Sensing: Cell-to-cell communication in Bacteria Annual Review of Cell and Development Biology 21: 319 - 346 | Waters, C. & Bassler, B. (2005) Quorum Sensing: Cell-to-cell communication in Bacteria Annual Review of Cell and Development Biology 21: 319 - 346 | ||
| − | </br> | + | </br></br> |
Yee Gyung Kwak, George A. Jacoby and David C. Hoopera. (2013) Induction of Plasmid-Carried qnrS1 in Escherichia coli by Naturally Occurring Quinolones and Quorum-Sensing Signal Molecules | Yee Gyung Kwak, George A. Jacoby and David C. Hoopera. (2013) Induction of Plasmid-Carried qnrS1 in Escherichia coli by Naturally Occurring Quinolones and Quorum-Sensing Signal Molecules | ||
Revision as of 02:20, 30 October 2017
spacefill
spacefill


























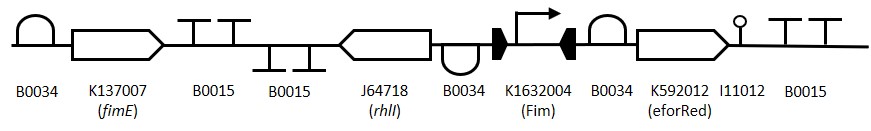
 Figure 2:
Representation of the switching mechanism of the Fim Switch, in the native [OFF] state the eforRED reporter is expressed (shown in red) allowing direct visualisation of the cells. Following the inversion of the promoter region (
Figure 2:
Representation of the switching mechanism of the Fim Switch, in the native [OFF] state the eforRED reporter is expressed (shown in red) allowing direct visualisation of the cells. Following the inversion of the promoter region ( Table 1: Table of parts used for constructing the Fim Switch.
Table 1: Table of parts used for constructing the Fim Switch. Figure 3:
High fidelity amplification of the 3 gBlock fragments for assembly of the Fim Switch. The gBlock-1 amplification is shown in lanes 1+2 (819 bp), gBlock-2 amplification is shown in lanes 3+4 (1148 bp) and the gBlock-3 amplification is shown in lanes 5+6 (939bp).
Figure 3:
High fidelity amplification of the 3 gBlock fragments for assembly of the Fim Switch. The gBlock-1 amplification is shown in lanes 1+2 (819 bp), gBlock-2 amplification is shown in lanes 3+4 (1148 bp) and the gBlock-3 amplification is shown in lanes 5+6 (939bp).
 Figure 4:
Patches of the Fim Switch transformants. Patch number 6 shows the correct red colour which indicates expression of the eforRed chromoprotein.
Figure 4:
Patches of the Fim Switch transformants. Patch number 6 shows the correct red colour which indicates expression of the eforRed chromoprotein.













 The
The 












































