| (31 intermediate revisions by 5 users not shown) | |||
| Line 110: | Line 110: | ||
.result_picture td p { | .result_picture td p { | ||
text-align: center; | text-align: center; | ||
| + | } | ||
| + | |||
| + | .keyres_hide { | ||
| + | display: none; | ||
| + | } | ||
| + | |||
| + | #key_res:hover { | ||
| + | cursor:pointer; | ||
} | } | ||
| Line 118: | Line 126: | ||
<div class="container-fluid text-center" style="max-width: 80%; font-size: 1em"> | <div class="container-fluid text-center" style="max-width: 80%; font-size: 1em"> | ||
| − | <h1 style=" | + | <h1 class="display 4" style="font-family: Rubik; margin: 0">Our Experimental Results</h1> |
<br /> | <br /> | ||
| + | <br /> | ||
| + | <div class="jumbotron rounded" style="background-color: #e8e8e8; border: 1px solid #222222; padding: 10px !important;"> | ||
| + | <h2 id="key_res" class="display-5 keyres_show" style="color: #222222">Key Achievements - click to show</h2> | ||
| + | <hr class="keyres_hide" style="color: #222222"> | ||
| + | <ul class="keyres_hide" style="color: #222222"> | ||
| + | |||
| + | |||
| + | <li class="keyres_hide" style="font-family: Rubik">Demonstrated that biosensors can be successfully split into three modules</li> | ||
| + | |||
| + | <li class="keyres_hide" style="font-family: Rubik">Produced biosensor variants by co-culturing different module variants together</li> | ||
| + | |||
| + | <li class="keyres_hide" style="font-family: Rubik">Used 3D spatial modelling to begin optimisation of a multicellular biosensor</li> | ||
| + | |||
| + | <li class="keyres_hide" style="font-family: Rubik">Characterised a 'standby switch' based on an improved part <a href="http://parts.igem.org/Part:BBa_K1632007">(BBa_K1632007)</a></li> | ||
| + | |||
| + | <li class="keyres_hide" style="font-family: Rubik">Demonstrated that a Design of Experiments approach can be used to optimise cell-free systems</li> | ||
| + | |||
| + | |||
| + | </ul> | ||
| + | </div> | ||
| + | |||
| + | |||
<p> | <p> | ||
Below is a diagram of our Sensynova Framework. Clicking on each part of the framework (e.g. detector modules) links to the relevant results.<br /> | Below is a diagram of our Sensynova Framework. Clicking on each part of the framework (e.g. detector modules) links to the relevant results.<br /> | ||
| Line 206: | Line 236: | ||
<center> | <center> | ||
<p>Multicellular Framework Testing</p> | <p>Multicellular Framework Testing</p> | ||
| − | |||
<a href="https://2017.igem.org/Team:Newcastle/Results#myTab"><img onClick="showTab('framework')" title="Full Framework in Cells" id="framework_res" class="hoverable hover_large" src="https://static.igem.org/mediawiki/2017/7/7b/T--Newcastle--BB_flask_results.png"/></a> | <a href="https://2017.igem.org/Team:Newcastle/Results#myTab"><img onClick="showTab('framework')" title="Full Framework in Cells" id="framework_res" class="hoverable hover_large" src="https://static.igem.org/mediawiki/2017/7/7b/T--Newcastle--BB_flask_results.png"/></a> | ||
</center> | </center> | ||
| Line 290: | Line 319: | ||
<center> | <center> | ||
<p>Framework in Cell Free Protein Synthesis Systems</p> | <p>Framework in Cell Free Protein Synthesis Systems</p> | ||
| − | |||
<a href="https://2017.igem.org/Team:Newcastle/Results#myTab"><img onClick="showTab('cellfree')" id="cellfree_res" title="Cell Free Protein Synthesis Systems" class="hoverable hover_large" src="https://static.igem.org/mediawiki/2017/b/b6/T--Newcastle--BB_CFPS_results.png"/></a> | <a href="https://2017.igem.org/Team:Newcastle/Results#myTab"><img onClick="showTab('cellfree')" id="cellfree_res" title="Cell Free Protein Synthesis Systems" class="hoverable hover_large" src="https://static.igem.org/mediawiki/2017/b/b6/T--Newcastle--BB_CFPS_results.png"/></a> | ||
</center> | </center> | ||
| Line 387: | Line 415: | ||
<h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Rationale and Aim </h2> | <h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Rationale and Aim </h2> | ||
</br> | </br> | ||
| − | <p>Glyphosate is a herbicide that works by blocking the activity of the enzyme enolpyruvylshikimate-3-phosphate synthase (EPSPS), which converts carbohydrates derived from glycolysis and the pentose phosphate pathway to plant metabolites and aromatic amino acids. | + | <p>Glyphosate is a herbicide that works by blocking the activity of the enzyme enolpyruvylshikimate-3-phosphate synthase (EPSPS), which converts carbohydrates derived from glycolysis and the pentose phosphate pathway to plant metabolites and aromatic amino acids. It was identified as a biosensor target through our <a href="https://2017.igem.org/Team:Newcastle/HP/Gold_Integrated">human practices</a> when we attended the N8 conference. |
</br></br> | </br></br> | ||
| − | We attempted to design a system capable of glyphosate detection. With little information regarding mechanisms of glyphosate interactions within the cell, we could not identify a simple system in which a responsive transcription factor was able to affect the production of a reporter gene. This is a common issue in many biosensor projects. | + | We attempted to design a system capable of glyphosate detection. With little information regarding mechanisms of glyphosate interactions within the cell, we could not identify a simple system in which a responsive transcription factor was able to affect the production of a reporter gene. This is a common issue in many biosensor projects. |
| + | To show the adaptor in action we chose to develop a part that would measure the level of glyphosate through the production of formaldehyde. There are known sensors for formaldehyde such as <a href="https://2012.igem.org/Team:TMU-Tokyo">Tokyo’s 2012 biosensor</a>. Our design relies on the natural biochemical systems, the c-p lyase pathways, in <i>E. coli</i> to convert glyphosate to sarcosine. We then designed a part, SOX, based on the production of the enzyme sarcosine oxidase, encoded by <i>soxA</i> to convert sarcosine to formaldehyde ready for detection by a formaldehyde producing input module. | ||
| + | |||
</br></br> | </br></br> | ||
The mining of transcriptome data has previously been used to find responsive DNA elements to a molecule of interest (Groningen 2012). Therefore, we analysed differences in transcriptome data between glyphosate sensitive and insensitive plants. A number of genes were found which were differently expressed. However, it was determined that it is more likely that this differential expression was not due to glyphosate directly, but rather the aromatic amino acid starvation caused by EPSPS inhibition by glyphosate, making these systems unsuitable for direct glyphosate detection. Various other systems we designed were also far from ideal, with high levels of complexity and reliance on native plant machinery. | The mining of transcriptome data has previously been used to find responsive DNA elements to a molecule of interest (Groningen 2012). Therefore, we analysed differences in transcriptome data between glyphosate sensitive and insensitive plants. A number of genes were found which were differently expressed. However, it was determined that it is more likely that this differential expression was not due to glyphosate directly, but rather the aromatic amino acid starvation caused by EPSPS inhibition by glyphosate, making these systems unsuitable for direct glyphosate detection. Various other systems we designed were also far from ideal, with high levels of complexity and reliance on native plant machinery. | ||
| Line 469: | Line 499: | ||
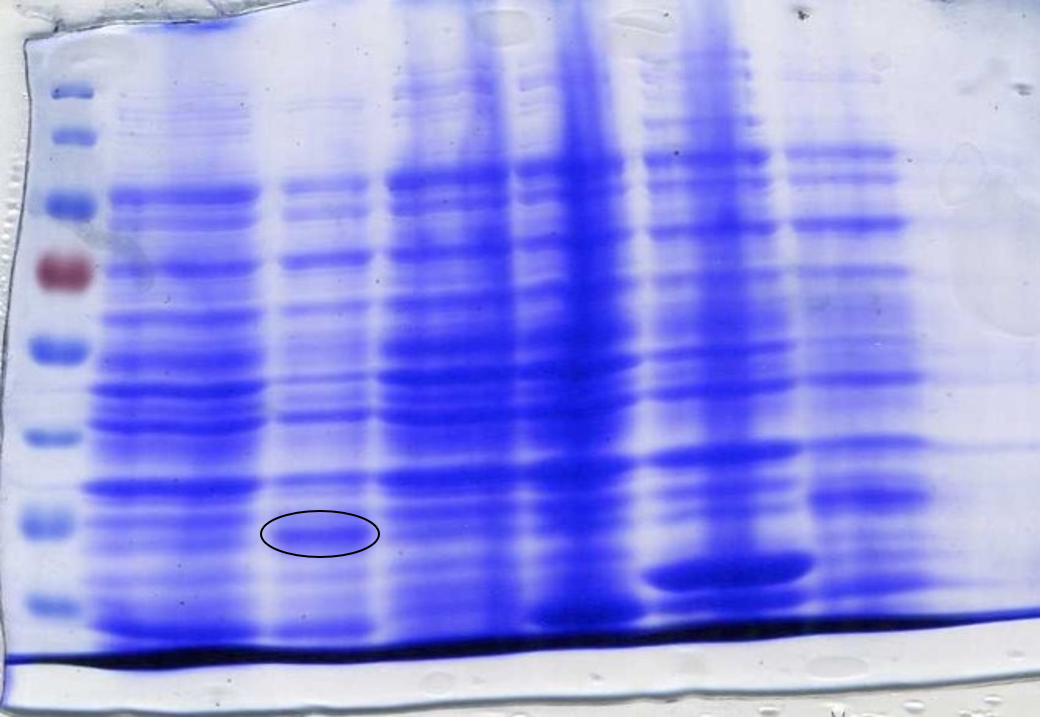
<p>To determine whether the now correct SOX had been successfully expressed another SDS-Page gel was performed. After inducing, harvesting and washing the cells 1 ml was taken from each culture to be loaded into the gel. The cells were lysed using lysozyme and boiled for 3 minutes at 100°C loading 10 µl into the gel (Figure 7). | <p>To determine whether the now correct SOX had been successfully expressed another SDS-Page gel was performed. After inducing, harvesting and washing the cells 1 ml was taken from each culture to be loaded into the gel. The cells were lysed using lysozyme and boiled for 3 minutes at 100°C loading 10 µl into the gel (Figure 7). | ||
</br></br> | </br></br> | ||
| − | <div class="SOX"><img src="https://static.igem.org/mediawiki/2017/8/89/T--Newcastle--Correct_sox_protein_gel_2.png" width="30%" style="background-color:white; margin-right: 2%; margin-bottom: 2%;" alt="" class="img-fluid border border-dark rounded mx-auto d-block"/> | + | <div class="SOX"><img src="https://static.igem.org/mediawiki/2017/8/89/T--Newcastle--Correct_sox_protein_gel_2.png" width="30%" style="background-color:white; margin-right: 2%; margin-bottom: 2%;" alt="" class="img-fluid border border-dark rounded mx-auto d-block"/> |
<br /> | <br /> | ||
<p class="legend"><center><strong>Figure 7:</strong> Sarcosine Oxidase expression was induced by adding 40 µl of 100 mM IPTG. Lane 1: 6 µl ladder, Lane 2: 10 µl sfGFP, Lane 3: BL21-DE3, Lane 4: 10µl SOX 1, Lane 5: 10 µl SOX 2, Lane 6: 10 µl SOX 3, Lane 7: 10 µl SOX 4, Lane 8: 10 µl SOX 5, Lane 9: 10 µl SOX 6, Lane 10: 6 µl ladder. Circled bands show sarcosine oxidase at ~42 kDa, the expected molecular weight.</p></center> | <p class="legend"><center><strong>Figure 7:</strong> Sarcosine Oxidase expression was induced by adding 40 µl of 100 mM IPTG. Lane 1: 6 µl ladder, Lane 2: 10 µl sfGFP, Lane 3: BL21-DE3, Lane 4: 10µl SOX 1, Lane 5: 10 µl SOX 2, Lane 6: 10 µl SOX 3, Lane 7: 10 µl SOX 4, Lane 8: 10 µl SOX 5, Lane 9: 10 µl SOX 6, Lane 10: 6 µl ladder. Circled bands show sarcosine oxidase at ~42 kDa, the expected molecular weight.</p></center> | ||
</div> | </div> | ||
</br></br> | </br></br> | ||
| − | + | <p>To test for the presence of formaldehyde, and to demonstrate this part works, larger cultures were grown following the aforementioned protocols, and the cells harvested, washed and lysed by sonication. 0 µl, 50 µl and 200 µl of Sarcosine at 0.9 g/50 ml concentration was added to the cell lysate and incubated at 37°C. Every 2.5 hours the lysate was tested for the presence of formaldehyde with commercial <a href="http://www.sigmaaldrich.com/catalog/product/sial/37072?lang=en®ion=GB">formaldehyde testing strips</a> (Figure 8).</p> | |
| − | </br></br> | + | </br></br> |
<div class="SOX"><img src="https://static.igem.org/mediawiki/2017/4/4b/T--Newcastle--SOX_testing.JPG" width="30%" style="background-color:white; margin-right: 2%; margin-bottom: 2%;" alt="" class="img-fluid border border-dark rounded mx-auto d-block"/> | <div class="SOX"><img src="https://static.igem.org/mediawiki/2017/4/4b/T--Newcastle--SOX_testing.JPG" width="30%" style="background-color:white; margin-right: 2%; margin-bottom: 2%;" alt="" class="img-fluid border border-dark rounded mx-auto d-block"/> | ||
<br /><br /> | <br /><br /> | ||
| Line 481: | Line 511: | ||
</div> | </div> | ||
</br></br> | </br></br> | ||
| − | <p> | + | <p>Formaldehyde was detected, showing that SOX works as expected, however there is slight leaky expression as formaldehyde is produced when no IPTG is added.</p> |
</p> | </p> | ||
| − | + | </br> | |
| − | + | <p> We also decided to add Glyphosate to determine the efficiency of the native C-P Lyase pathway. Everything was repeated the same but instead we added 0 µl, 20 µl, 200 µl and 2 ml of glyphosate at 10mg/L. After 8 hours of testing and left overnight, none of the samples had produced formaldehyde according to the testing strips. The testing strips detect a minimum formaldehyde concentration of 10 mg/L, so it was possible that formaldehyde had been produced but that there was too little of it to detect with the strips.</p> | |
| − | + | ||
| − | + | </br> | |
| − | + | ||
| − | + | ||
<h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Conclusions and Future Work </h2> | <h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Conclusions and Future Work </h2> | ||
</br> | </br> | ||
| − | <p><i>E. coli</i> cells naturally have the C-P lyase pathway which degrades glyphosate into sarcosine. The fact that | + | <p><i>E. coli</i> cells naturally have the C-P lyase pathway which degrades glyphosate into sarcosine. The fact that formaldehyde was produced when sarcosine was added, but not when glyphosate was added, indicates that we have not overexpressed the C-P lyase pathway enough to produce enough sarcosine for SOX to convert into formaldehyde to be detected. |
</br></br> | </br></br> | ||
<p>Due to time constraints, we were unable to produce an <i>in vivo</i> formaldehyde detector variant of the Sensynova framework. Future characterisation of this part would include using the platform customised as a formaldehyde biosensor in order to sense compound produce and therefore creating a biosensor of glyphosate. | <p>Due to time constraints, we were unable to produce an <i>in vivo</i> formaldehyde detector variant of the Sensynova framework. Future characterisation of this part would include using the platform customised as a formaldehyde biosensor in order to sense compound produce and therefore creating a biosensor of glyphosate. | ||
| Line 539: | Line 567: | ||
<center><b>Figure 2:</b> Graph Indicating the Most Frequent Spacer Between -35 and -10 Regions Found in <i>E. coli</i> Promoters. This image was taken from Harley and Reynolds (1987).</center> | <center><b>Figure 2:</b> Graph Indicating the Most Frequent Spacer Between -35 and -10 Regions Found in <i>E. coli</i> Promoters. This image was taken from Harley and Reynolds (1987).</center> | ||
</p> | </p> | ||
| − | </br> | + | </br> |
<h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Design Stage </h2> | <h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Design Stage </h2> | ||
</br> | </br> | ||
| Line 585: | Line 613: | ||
<h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Conclusions and Future Work </h2> | <h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Conclusions and Future Work </h2> | ||
</br> | </br> | ||
| − | <p>Though we have generated a sizable library of promoters of varying strengths and functions, we lacked the time to complete its characterization by the screening against targeted molecules. | + | <p>Though we have generated a sizable library of promoters of varying strengths and functions, we lacked the time to complete its characterization by the screening against targeted molecules. |
<br/><br/> | <br/><br/> | ||
Due to time constraints, we also lacked the time to characterise these parts into the Sensynova platform within the lab. | Due to time constraints, we also lacked the time to characterise these parts into the Sensynova platform within the lab. | ||
| Line 592: | Line 620: | ||
<h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> References </h2> | <h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> References </h2> | ||
</br> | </br> | ||
| − | <p>Becker, N., Peters, J., Lionberger, T. and Maher, L. (2012). Mechanism of promoter repression by Lac repressor–DNA loops. Nucleic Acids Research, 41(1), pp.156-166. <br/> | + | <p>Becker, N., Peters, J., Lionberger, T. and Maher, L. (2012). Mechanism of promoter repression by Lac repressor–DNA loops. Nucleic Acids Research, 41(1), pp.156-166. <br/><br/> |
DeBoer, H. (1985). Microbial hybrid promoters. US4551433 A. | DeBoer, H. (1985). Microbial hybrid promoters. US4551433 A. | ||
<br/><br/> | <br/><br/> | ||
| Line 628: | Line 656: | ||
<img src="https://static.igem.org/mediawiki/2017/b/b1/Vava1aa.png" class="img-fluid border border-dark rounded" style="margin: 2%; max-width: 70%"> | <img src="https://static.igem.org/mediawiki/2017/b/b1/Vava1aa.png" class="img-fluid border border-dark rounded" style="margin: 2%; max-width: 70%"> | ||
| − | <center><b>Figure 1:</b> | + | <center><b>Figure 1:</b> <a href="http://parts.igem.org/Part:BBa_J33201">BBa_J33201</a> design.</center></p> |
</br> | </br> | ||
| Line 641: | Line 669: | ||
<td> | <td> | ||
<img src="https://static.igem.org/mediawiki/2017/d/db/Vave2.jpg" class="img-fluid border border-dark rounded" style="margin: 2%; max-width: 70%"> | <img src="https://static.igem.org/mediawiki/2017/d/db/Vave2.jpg" class="img-fluid border border-dark rounded" style="margin: 2%; max-width: 70%"> | ||
| − | <div><p class="legend"><center><strong><b>Figure 2:</b> </strong> | + | <div><p class="legend"><center><strong><b>Figure 2:</b> </strong> The new part <a href="http://parts.igem.org/Part:BBa_K2205022">BBa_K2205022</a> based on the previous Arsenic detector design <a href="http://parts.igem.org/Part:BBa_J33201">BBa_J33201</a>, implemented by the Sensynova part coding for the connector 1, <a href="http://parts.igem.org/Part:BBa_K2205008">BBa_K2205008</a>.</p></center></div> |
</td> | </td> | ||
| Line 716: | Line 744: | ||
<h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> References </h2> | <h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> References </h2> | ||
</br> | </br> | ||
| − | <p>Brenner K, Karing D, Weiss R, Arnold F (2007) Engineered bidirectional communication mediates a consensus in a microbial biofilm consortium. Proc Natl Acad Sci USA 104(44): 17300 - 17304 </br> de Mora K, Joshi N, Balint BL, Ward FB, Elfick A, French CE (2011) A pH-based biosensor for detection of arsenic in drinking water. Anal Bioanal Chem 400(4):1031-9 (Epub 2011 Mar 27).</p> | + | <p>Brenner K, Karing D, Weiss R, Arnold F (2007) Engineered bidirectional communication mediates a consensus in a microbial biofilm consortium. Proc Natl Acad Sci USA 104(44): 17300 - 17304 <br/><br/> de Mora K, Joshi N, Balint BL, Ward FB, Elfick A, French CE (2011) A pH-based biosensor for detection of arsenic in drinking water. Anal Bioanal Chem 400(4):1031-9 (Epub 2011 Mar 27).</p> |
</div> | </div> | ||
| Line 791: | Line 819: | ||
<h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Characterisation</h2> | <h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Characterisation</h2> | ||
</br> | </br> | ||
| − | <p> A preliminary qualitative assay was carried out as an initial test for this construct. Co-cultures of Psicose detector, processor unit and sfGFP reporter (<a href="http://parts.igem.org/wiki/index.php?title=Part:BBa_K2205015">BBa_K2205015</a>) were inoculated and grown overnight in LB+chloramphenicol (12.5 ng/ul). | + | <p> A preliminary qualitative assay was carried out as an initial test for this construct. Co-cultures of Psicose detector, processor unit and sfGFP reporter (<a href="http://parts.igem.org/wiki/index.php?title=Part:BBa_K2205015">BBa_K2205015</a>) were inoculated and grown overnight in LB+chloramphenicol (12.5 ng/ul). |
</br></br> | </br></br> | ||
The day after the cultures were diluted at OD600 0.1 and mixed together to obtain co-cultures with ratio 1:1:13 (detector:processor:reporter). The samples were supplemented with 33.22 mM Psicose to induce the expression of quorum sensing molecules and eventually achieve the reporter visualisation (Figures 8). </p> | The day after the cultures were diluted at OD600 0.1 and mixed together to obtain co-cultures with ratio 1:1:13 (detector:processor:reporter). The samples were supplemented with 33.22 mM Psicose to induce the expression of quorum sensing molecules and eventually achieve the reporter visualisation (Figures 8). </p> | ||
| Line 815: | Line 843: | ||
<hr> | <hr> | ||
| − | + | ||
<h1 style="font-family: Rubik"> Formaldehyde <button class="btn btn-primary collapsed" type="button" data-toggle="collapse" data-target="#formaldehyde" aria-expanded="false" aria-controls="formaldehyde" style="margin-left: 1%"></button></h1> | <h1 style="font-family: Rubik"> Formaldehyde <button class="btn btn-primary collapsed" type="button" data-toggle="collapse" data-target="#formaldehyde" aria-expanded="false" aria-controls="formaldehyde" style="margin-left: 1%"></button></h1> | ||
<div id="formaldehyde" class="collapse"> | <div id="formaldehyde" class="collapse"> | ||
| Line 821: | Line 849: | ||
<h2 style="font-size: 1em"> BioBricks used: <a href="http://parts.igem.org/Part:BBa_K2205029">BBa_K2205029 (New)</a>, <a href="http://parts.igem.org/Part:BBa_K2205030">BBa_K2205030 (New)</a>, <a href="http://parts.igem.org/Part:BBa_K749021">BBa_K749021(TMU-Tokyo 2012 )</a> </h2> | <h2 style="font-size: 1em"> BioBricks used: <a href="http://parts.igem.org/Part:BBa_K2205029">BBa_K2205029 (New)</a>, <a href="http://parts.igem.org/Part:BBa_K2205030">BBa_K2205030 (New)</a>, <a href="http://parts.igem.org/Part:BBa_K749021">BBa_K749021(TMU-Tokyo 2012 )</a> </h2> | ||
</br> | </br> | ||
| − | + | ||
<h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Rationale and Aim </h2> | <h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Rationale and Aim </h2> | ||
</br> | </br> | ||
| Line 832: | Line 860: | ||
<h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Background Information </h2> | <h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Background Information </h2> | ||
</br> | </br> | ||
| − | <p>The formaldehyde biosensor, part BBa_K749021, was selected was originally made and submitted to the iGEM registry by the TMU-Tokyo 2012 team. | + | <p>The formaldehyde biosensor, part BBa_K749021, was selected was originally made and submitted to the iGEM registry by the TMU-Tokyo 2012 team. |
</br></br> | </br></br> | ||
| − | This part was chosen as a variant to the detector module present in the Sensynova platform due to the fact that our adaptor module present in the framework, Sarcosine Oxidase, was made in order to convert glyphosate into formaldehyde, in order to overcome the limitation in the detection of glyphosate due to its little-known knowledge. | + | This part was chosen as a variant to the detector module present in the Sensynova platform due to the fact that our adaptor module present in the framework, Sarcosine Oxidase, was made in order to convert glyphosate into formaldehyde, in order to overcome the limitation in the detection of glyphosate due to its little-known knowledge. |
</br> | </br> | ||
<center><img src="https://static.igem.org/mediawiki/2017/1/1f/T--Newcastle--Lais--FO--Ruler.png" class="img-fluid border border-dark rounded" style="margin: 2%"></center> | <center><img src="https://static.igem.org/mediawiki/2017/1/1f/T--Newcastle--Lais--FO--Ruler.png" class="img-fluid border border-dark rounded" style="margin: 2%"></center> | ||
| Line 853: | Line 881: | ||
<h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Design Stage </h2> | <h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Design Stage </h2> | ||
</br> | </br> | ||
| − | <p>In order to implement the Formaldehyde biosensor variant to the Sensynova platform, a design was created by replacing the IPTG sensing system in the original detector module with the construct detailed above, creating part <a href="http://parts.igem.org/Part:BBa_K2205030">BBa_K2205030 </a>. | + | <p>In order to implement the Formaldehyde biosensor variant to the Sensynova platform, a design was created by replacing the IPTG sensing system in the original detector module with the construct detailed above, creating part <a href="http://parts.igem.org/Part:BBa_K2205030">BBa_K2205030 </a>. |
</br></br> | </br></br> | ||
We chose to redesign the Formaldehyde biosensor detailed above to mirror the design used when producing the Psicose detector variant. The system detailed in the image below is made up of the constitutive promoter present within the platform triggering transcription of the FrmR repressing the PfrmR and subsequently the connector 1 of the Sensynova platform. We have also replaced the colour output present in the TMU-Tokyo design, we have added our part <a href="http://parts.igem.org/Part:BBa_K2205008">BBa_K2205008</a>, which produces our first connector in order to trigger a response from following modules of the Sensynova platform in the presence of Formaldehyde.</p> | We chose to redesign the Formaldehyde biosensor detailed above to mirror the design used when producing the Psicose detector variant. The system detailed in the image below is made up of the constitutive promoter present within the platform triggering transcription of the FrmR repressing the PfrmR and subsequently the connector 1 of the Sensynova platform. We have also replaced the colour output present in the TMU-Tokyo design, we have added our part <a href="http://parts.igem.org/Part:BBa_K2205008">BBa_K2205008</a>, which produces our first connector in order to trigger a response from following modules of the Sensynova platform in the presence of Formaldehyde.</p> | ||
| Line 883: | Line 911: | ||
<h2 style="font-family: Rubik; text-align: left; margin-top: 1%">Conclusions and Future Work </h2> | <h2 style="font-family: Rubik; text-align: left; margin-top: 1%">Conclusions and Future Work </h2> | ||
</br> | </br> | ||
| − | <p>Due to time constraints, we lacked the time to synthesise, implement and characterise this part into the Sensynova platform within the lab. Future work on this part would include characterisation <i>in vivo</i> guided by the modelling of the framework when customised as a formaldehyde biosensor and testing against the Sarcosine Oxidase adaptor module currently present in the framework. | + | <p>Due to time constraints, we lacked the time to synthesise, implement and characterise this part into the Sensynova platform within the lab. Future work on this part would include characterisation <i>in vivo</i> guided by the modelling of the framework when customised as a formaldehyde biosensor and testing against the Sarcosine Oxidase adaptor module currently present in the framework. |
</p> | </p> | ||
</br> | </br> | ||
| Line 894: | Line 922: | ||
</div> | </div> | ||
</div> | </div> | ||
| − | + | ||
<div class="tab-pane fade" id="nav-processor" role="tabpanel" aria-labelledby="nav-processor-tab"> | <div class="tab-pane fade" id="nav-processor" role="tabpanel" aria-labelledby="nav-processor-tab"> | ||
</br> | </br> | ||
| Line 1,232: | Line 1,260: | ||
</br> | </br> | ||
<p> | <p> | ||
| − | Li X., Zhang G., Ngo N., Zhao X., Kain S.R., Huang C.C., (1997), Deletions of the Aequorea victoria green fluorescent protein define the minimal domain required for fluorescence, <i>J. Biol. Chem.</i>, 272:28545–9, doi: 10.1074/jbc.272.45.28545<br /> | + | Li X., Zhang G., Ngo N., Zhao X., Kain S.R., Huang C.C., (1997), Deletions of the Aequorea victoria green fluorescent protein define the minimal domain required for fluorescence, <i>J. Biol. Chem.</i>, 272:28545–9, doi: 10.1074/jbc.272.45.28545 <br/><br/> |
Shin, J., and Noireaux, V., (2010), Efficient Cell-Free Expression with the Endogenous <i>E. coli</i> RNA Polymerase and Sigma Factor 70, <i>J. Biol. Eng.</i>, 4:8, doi: 10.1186/1754-1611-4-8<br /> | Shin, J., and Noireaux, V., (2010), Efficient Cell-Free Expression with the Endogenous <i>E. coli</i> RNA Polymerase and Sigma Factor 70, <i>J. Biol. Eng.</i>, 4:8, doi: 10.1186/1754-1611-4-8<br /> | ||
| Line 1,705: | Line 1,733: | ||
<div> | <div> | ||
| − | <img src="https://static.igem.org/mediawiki/2017/1/1d/T--Newcastle--BB_pSB1C3-sfGFP_plasmid_map.png" width="400px" class="img-fluid border border-dark rounded mx-auto d-block" style="background-color:white; margin-right: 2%; margin-bottom: 2%; alt=""/> | + | <img src="https://static.igem.org/mediawiki/2017/1/1d/T--Newcastle--BB_pSB1C3-sfGFP_plasmid_map.png" width="400px" class="img-fluid border border-dark rounded mx-auto d-block" style="background-color:white; margin-right: 2%; margin-bottom: 2%;" alt=""/> |
<p class="legend"><center><strong>Figure 2:</strong> Plasmid map for pSB1C3-sfGFP. Construct is standard biobrick part BBa_ K515105.</center></p> | <p class="legend"><center><strong>Figure 2:</strong> Plasmid map for pSB1C3-sfGFP. Construct is standard biobrick part BBa_ K515105.</center></p> | ||
</div> | </div> | ||
| Line 1,760: | Line 1,788: | ||
<div> | <div> | ||
| − | <img src="https://static.igem.org/mediawiki/2017/b/b0/T--Newcastle--BB_CFPS_figure4.png" width="600px" class="img-fluid border border-dark rounded mx-auto d-block" style="background-color:white; margin-right: 2%; margin-bottom: 2%; alt=""/> | + | <img src="https://static.igem.org/mediawiki/2017/b/b0/T--Newcastle--BB_CFPS_figure4.png" width="600px" class="img-fluid border border-dark rounded mx-auto d-block" style="background-color:white; margin-right: 2%; margin-bottom: 2%;" alt=""/> |
<p class="legend"><center><strong>Figure 5:</strong> Screening model constructed using JMP showing which factors were closest to significance. Predictions for interactions are unreliable due to forced orthogonality (*). Of the primary factors, magnesium glutamate is the closest to significant, followed by potassium glutamate, sodium oxalate, and ammonium acetate in that order.</center></p> | <p class="legend"><center><strong>Figure 5:</strong> Screening model constructed using JMP showing which factors were closest to significance. Predictions for interactions are unreliable due to forced orthogonality (*). Of the primary factors, magnesium glutamate is the closest to significant, followed by potassium glutamate, sodium oxalate, and ammonium acetate in that order.</center></p> | ||
</div> | </div> | ||
| Line 1,878: | Line 1,906: | ||
</div> | </div> | ||
</br> | </br> | ||
| + | |||
| + | |||
| + | |||
<h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Conclusions and Future Work </h2> | <h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> Conclusions and Future Work </h2> | ||
| + | </br> | ||
<p>This study has begun multifactorial analysis on the components of the supplemental solution for cell free protein synthesis systems. It has provided evidence that some supplements have a greater effect on a systems protein synthesis activity than others, and that the important factors may differ between cell extract batches. The ability to use a Design of Experiments approach towards the optimisation of CFPS systems has also been demonstrated. While this study has provided evidence towards these claims, further work should be performed to validate the findings. A DoE screening design for the supplements of CFPS systems should be used on the same cell extract batch repeatedly. This will help confirm that the screening models derived from the experimental design data are accurate. The screening design should also be performed on many different batches of at least moderately active cell extracts to confirm that important supplements do differ between batches. | <p>This study has begun multifactorial analysis on the components of the supplemental solution for cell free protein synthesis systems. It has provided evidence that some supplements have a greater effect on a systems protein synthesis activity than others, and that the important factors may differ between cell extract batches. The ability to use a Design of Experiments approach towards the optimisation of CFPS systems has also been demonstrated. While this study has provided evidence towards these claims, further work should be performed to validate the findings. A DoE screening design for the supplements of CFPS systems should be used on the same cell extract batch repeatedly. This will help confirm that the screening models derived from the experimental design data are accurate. The screening design should also be performed on many different batches of at least moderately active cell extracts to confirm that important supplements do differ between batches. | ||
</br></br> | </br></br> | ||
| − | Following the above, a surface response design could be used for all commonly important supplements of the CFPS system to determine its effectiveness at optimising CFPS activity. The information could also be used to determine commonly unimportant supplements so they can be eliminated from the supplement solution, hence decreasing the cost per reaction.</p> | + | Following the above, a surface response design could be used for all commonly important supplements of the CFPS system to determine its effectiveness at optimising CFPS activity. The information could also be used to determine commonly unimportant supplements so they can be eliminated from the supplement solution, hence decreasing the cost per reaction. Furthermore, the full Sensynova system can be expressed in cell-free.</p> |
</br> | </br> | ||
<h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> References </h2> | <h2 style="font-family: Rubik; text-align: left; margin-top: 1%"> References </h2> | ||
| + | </br> | ||
<p> | <p> | ||
| − | Algranati, I. D. & Goldemberg, S. H., 1977. Polyamines and their role in protein synthesis. <i>Trends in Biochem. Sci.</i>, 2(12), pp. 272-274.<br /> | + | Algranati, I. D. & Goldemberg, S. H., 1977. Polyamines and their role in protein synthesis. <i>Trends in Biochem. Sci.</i>, 2(12), pp. 272-274. <br/><br/> |
| − | Anderson, M. J. & Whitcomb, P. J., 2010. Design of Experiments. In: Kirk-Othmer Encyclopedia of Chemical Technology. <i>s.l.:John Wiley & Sons, Inc</i>, pp. 1-22. <br /> | + | Anderson, M. J. & Whitcomb, P. J., 2010. Design of Experiments. In: Kirk-Othmer Encyclopedia of Chemical Technology. <i>s.l.:John Wiley & Sons, Inc</i>, pp. 1-22. <br/><br/> |
| − | Borg, A. & Ehrenberg, M., 2015. Determinants of the Rate of mRNA Translocation in Bacterial Protein Synthesis. <i>J. Mol. Biol.</i>, 427(9), pp. 1835-1847.<br /> | + | Borg, A. & Ehrenberg, M., 2015. Determinants of the Rate of mRNA Translocation in Bacterial Protein Synthesis. <i>J. Mol. Biol.</i>, 427(9), pp. 1835-1847. <br/><br/> |
| − | Carlson, E. D., Gan, R., Hodgman, C. E. & Jewett, M. C., 2012. Cell-Free Protein Synthesis: Applications Come of Age. <i>Biotechnol. Adv.</i>, 30(5), pp. 1185-1194.<br /> | + | Carlson, E. D., Gan, R., Hodgman, C. E. & Jewett, M. C., 2012. Cell-Free Protein Synthesis: Applications Come of Age. <i>Biotechnol. Adv.</i>, 30(5), pp. 1185-1194. <br/><br/> |
| − | Garamella, J., Marshall, R., Rustad, M. & Noireaux, V., 2016. The All E. coli TX-TL Toolbox 2.0: A Platform for Cell-Free Synthetic Biology. <i>ACS Syn. Biol.</i>, 5(4), pp. 344-355.<br /> | + | Garamella, J., Marshall, R., Rustad, M. & Noireaux, V., 2016. The All E. coli TX-TL Toolbox 2.0: A Platform for Cell-Free Synthetic Biology. <i>ACS Syn. Biol.</i>, 5(4), pp. 344-355. <br/><br/> |
| − | Jelenc, P. C. & Kurland, C. G., 1979. Nucleoside triphosphate regeneration decreases the frequency of translation errors. <i>Proc. Natl. Acad. Sci. USA</i>, 76(7), pp. 3174-3178.<br /> | + | Jelenc, P. C. & Kurland, C. G., 1979. Nucleoside triphosphate regeneration decreases the frequency of translation errors. <i>Proc. Natl. Acad. Sci. USA</i>, 76(7), pp. 3174-3178. <br/><br/> |
| − | Jewett, M. C. & Swartz, J. R., 2004. Mimicking the <i>Escherichia coli</i> cytoplasmic environment activates long-lived and efficient cell-free protein synthesis. <i>Biotechnol. & Bioeng.</i>, 86(1), pp. 19-26.<br /> | + | Jewett, M. C. & Swartz, J. R., 2004. Mimicking the <i>Escherichia coli</i> cytoplasmic environment activates long-lived and efficient cell-free protein synthesis. <i>Biotechnol. & Bioeng.</i>, 86(1), pp. 19-26. <br/><br/> |
| − | Jewett, M. C. et al., 2008. An integrated cell-free metabolic platform for protein production and synthetic biology. <i>Mol. Syst. Biol.</i>, 4(220).<br /> | + | Jewett, M. C. et al., 2008. An integrated cell-free metabolic platform for protein production and synthetic biology. <i>Mol. Syst. Biol.</i>, 4(220). <br/><br/> |
| − | Katsura, K. et al., 2017. A reproducible and scalable procedure for preparing bacterial extracts for cell-free protein synthesis. <i>J. Biochem.</i>, 162(5), pp. 357-369.<br /> | + | Katsura, K. et al., 2017. A reproducible and scalable procedure for preparing bacterial extracts for cell-free protein synthesis. <i>J. Biochem.</i>, 162(5), pp. 357-369. <br/><br/> |
| − | Kelwick, R., Webb, A. J., MacDonald, J. & Freemont, P. S., 2016. Development of a Bacillus subtilis cell-free transcription-translation system for prototyping regulatory elements. <i>Metab. Eng.</i>, Volume 38, pp. 370-381.<br /> | + | Kelwick, R., Webb, A. J., MacDonald, J. & Freemont, P. S., 2016. Development of a Bacillus subtilis cell-free transcription-translation system for prototyping regulatory elements. <i>Metab. Eng.</i>, Volume 38, pp. 370-381. <br/><br/> |
| − | Kwon, Y. & Jewett, M. C., 2015. High-throughput preparation methods of crude extract for robust cell-free protein synthesis. <i>Sci. Rep.</i>, Volume 5.<br /> | + | Kwon, Y. & Jewett, M. C., 2015. High-throughput preparation methods of crude extract for robust cell-free protein synthesis. <i>Sci. Rep.</i>, Volume 5. <br/><br/> |
| − | Lee, K. H. & Kim, D. M., 2013. Applications of cell-free protein synthesis in synthetic biology: Interfacing bio-machinery with synthetic environments. <i>Biotechnol. J.</i>, 8(11), pp. 1292-1300.<br /> | + | Lee, K. H. & Kim, D. M., 2013. Applications of cell-free protein synthesis in synthetic biology: Interfacing bio-machinery with synthetic environments. <i>Biotechnol. J.</i>, 8(11), pp. 1292-1300. <br/><br/> |
| − | Lu, Y., 2017. Cell-free synthetic biology: Engineering in an open world. <i>Syn. and Sys. Biotech.</i>, 2(1), pp. 23-27.<br /> | + | Lu, Y., 2017. Cell-free synthetic biology: Engineering in an open world. <i>Syn. and Sys. Biotech.</i>, 2(1), pp. 23-27. <br/><br/> |
| − | Nierhaus, K. H., 2014. Mg2+, K+, and the Ribosome. <i>J. Bacteriol.</i>, 196(22), pp. 3817-3819.<br /> | + | Nierhaus, K. H., 2014. Mg2+, K+, and the Ribosome. <i>J. Bacteriol.</i>, 196(22), pp. 3817-3819. <br/><br/> |
| − | Nirenberg, M. W. & Matthaei, J. H., 1961. The dependence of cell-free protein synthesis in <i>E. coli</i> upon naturally occurring or synthetic polyribonucleotides. <i>Proc. Natl. Acad. Sci. USA</i>, 47(10), pp. 1588-1602.<br /> | + | Nirenberg, M. W. & Matthaei, J. H., 1961. The dependence of cell-free protein synthesis in <i>E. coli</i> upon naturally occurring or synthetic polyribonucleotides. <i>Proc. Natl. Acad. Sci. USA</i>, 47(10), pp. 1588-1602. <br/><br/> |
| − | Pyle, A. M., 2002. Metal ions in the structure and function of RNA. <i>J. Biol. Inorg.</i>, 7(8), pp. 679-690.<br /> | + | Pyle, A. M., 2002. Metal ions in the structure and function of RNA. <i>J. Biol. Inorg.</i>, 7(8), pp. 679-690. <br/><br/> |
| − | SAS Institute Inc., 2016. JMP® 13 Design of Experiments Guide. Cary, NC, USA: SAS Institute Inc.<br /> | + | SAS Institute Inc., 2016. JMP® 13 Design of Experiments Guide. Cary, NC, USA: SAS Institute Inc. <br/><br/> |
Yang, W. C., Patel, K. & Wong, H. E., 2012. Simplifying and streamlining <i>Escherichia coli</i>-based cell-free protein synthesis. <i>Biotechnol. Prog.</i>, 28(2), pp. 413-420.<br /> | Yang, W. C., Patel, K. & Wong, H. E., 2012. Simplifying and streamlining <i>Escherichia coli</i>-based cell-free protein synthesis. <i>Biotechnol. Prog.</i>, 28(2), pp. 413-420.<br /> | ||
| Line 1,929: | Line 1,962: | ||
</div> | </div> | ||
</div> | </div> | ||
| − | + | ||
| + | |||
</div> | </div> | ||
| Line 1,935: | Line 1,969: | ||
</div> | </div> | ||
| + | </div> | ||
| Line 1,993: | Line 2,028: | ||
document.getElementById("nav-cellfree").classList.remove("active", "show"); | document.getElementById("nav-cellfree").classList.remove("active", "show"); | ||
} | } | ||
| + | |||
| + | |||
| + | $("#key_res").click(function(){ | ||
| + | $(".keyres_hide").toggle(); | ||
| + | }); | ||
Latest revision as of 22:39, 1 November 2017
spacefill
spacefill
Our Experimental Results
Key Achievements - click to show
- Demonstrated that biosensors can be successfully split into three modules
- Produced biosensor variants by co-culturing different module variants together
- Used 3D spatial modelling to begin optimisation of a multicellular biosensor
- Characterised a 'standby switch' based on an improved part (BBa_K1632007)
- Demonstrated that a Design of Experiments approach can be used to optimise cell-free systems
Below is a diagram of our Sensynova Framework. Clicking on each part of the framework (e.g. detector modules) links to the relevant results.
Alternatively, at the bottom of this page are tabs which will show you results for every part of the project
 |
 |
 |
 |
 |
 |
 |
 |






























































 The
The 











































