Cccxyyyyyyyy (Talk | contribs) |
|||
| (23 intermediate revisions by 5 users not shown) | |||
| Line 254: | Line 254: | ||
<div id="title"> | <div id="title"> | ||
<h1>Demonstrate</h1> | <h1>Demonstrate</h1> | ||
| − | <hr> | + | <a title="huge suprise" href="https://2017.igem.org/Team:Tianjin/surprise23333" target="_blank"><hr></a> |
</div> | </div> | ||
| Line 272: | Line 272: | ||
<h4>Obtaining the chassis </h4> | <h4>Obtaining the chassis </h4> | ||
<hr> | <hr> | ||
| − | <p>Aiming to achieve MTS for environmental use, it is essential to make sure that when the <i>MAT</i> locus has DSB (double strands break) cleaved by <i>HO</i>, our type-a (<i>MATa</i>) yeast can only become type-α (<i>MATα</i>). Therefore, we used a <i>Ura-tag</i> to replace the<i> HMRa</i> domain in <i>chromosome Ⅲ</i>. In this way the <i>HMRa</i> will no longer be the donor for the homologous recombination in the repairing process for MAT cleavage | + | <p>Aiming to achieve MTS for environmental use, it is essential to make sure that when the <i>MAT</i> locus has DSB (double strands break) cleaved by <i>HO</i>, our type-a (<i>MATa</i>) yeast can only become type-α (<i>MATα</i>). Therefore, we used a <i>Ura-tag</i> to replace the<i> HMRa</i> domain in <i>chromosome Ⅲ</i>. In this way the <i>HMRa</i> will no longer be the donor for the homologous recombination in the repairing process for MAT cleavage. After selection, by homologous recombination, we deleted the <i>Ura-tag</i> for further usage. We selected the target colonies (<b><i>SynⅩ-dUra</i></b>) via <i>5-FOA</i> plates. </p> |
<div class="zxx_zoom_demo"> | <div class="zxx_zoom_demo"> | ||
<div class="small_pic_demo" style="float:left;"> | <div class="small_pic_demo" style="float:left;"> | ||
| Line 282: | Line 282: | ||
<a href="#pic_eleven" > | <a href="#pic_eleven" > | ||
<img src="https://static.igem.org/mediawiki/2017/0/03/Tianjin-ho-result-9.jpeg"/> | <img src="https://static.igem.org/mediawiki/2017/0/03/Tianjin-ho-result-9.jpeg"/> | ||
| − | </a> <p style="font-size:15px;text-align:center"><br/>Fig. 1-2. As we can see in the gel photo above, the <b>UP</b> and <b>DOWN</b> segments hasn’t been amplified in our <i><b>SynⅩ-dUra</b></i> comparing to the BY4741 as control | + | </a> <p style="font-size:15px;text-align:center"><br/>Fig. 1-2. As we can see in the gel photo above, the <b>UP</b> and <b>DOWN</b> segments hasn’t been amplified in our <i><b>SynⅩ-dUra</b></i> comparing to the BY4741 as control, which indicated that the HMRa gene has been successfully eliminated. |
</p> | </p> | ||
</div> | </div> | ||
| Line 289: | Line 289: | ||
<div id="pic_ten" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/1/1b/Tianjin-ho-result-10.jpeg"/><p style="font-size:15px;text-align:center"><br/> Fig. 1-1. The PCR strategy for testing whether we deleted the HMRa in <b><i>SynⅩ-dUra</i></b>. | <div id="pic_ten" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/1/1b/Tianjin-ho-result-10.jpeg"/><p style="font-size:15px;text-align:center"><br/> Fig. 1-1. The PCR strategy for testing whether we deleted the HMRa in <b><i>SynⅩ-dUra</i></b>. | ||
</p></div> | </p></div> | ||
| − | <div id="pic_eleven" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/0/03/Tianjin-ho-result-9.jpeg"/><p style="font-size:15px;text-align:center"><br/>Fig. 1-2. As we can see in the gel photo above, the <b>UP</b> and <b>DOWN</b> segments hasn’t been amplified in our <i><b>SynⅩ-dUra</b></i> comparing to the BY4741 as control | + | <div id="pic_eleven" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/0/03/Tianjin-ho-result-9.jpeg"/><p style="font-size:15px;text-align:center"><br/>Fig. 1-2. As we can see in the gel photo above, the <b>UP</b> and <b>DOWN</b> segments hasn’t been amplified in our <i><b>SynⅩ-dUra</b></i> comparing to the BY4741 as control, which indicated that the HMRa gene has been successfully eliminated. |
</p></div> | </p></div> | ||
<h4> The result for constructing the <i>Gal</i> systems</h4> | <h4> The result for constructing the <i>Gal</i> systems</h4> | ||
| Line 328: | Line 328: | ||
<p> Cultivate two groups of yeasts together. (one is <b><i>SynⅩ-dUra-416</i></b>, the other is normal <i>BY4741 MATa</i>) If the <b>MTS</b> has been accomplished (<b><i>SynⅩ-dUra-416</i></b> can become <i>MATα</i>), the two groups of haploids can mate with each other and become diploids. </p> | <p> Cultivate two groups of yeasts together. (one is <b><i>SynⅩ-dUra-416</i></b>, the other is normal <i>BY4741 MATa</i>) If the <b>MTS</b> has been accomplished (<b><i>SynⅩ-dUra-416</i></b> can become <i>MATα</i>), the two groups of haploids can mate with each other and become diploids. </p> | ||
<h5>3) Step three</h5> | <h5>3) Step three</h5> | ||
| − | <p>Test the results of mating by PCR method. We designed the primers for both <i>MATa</i> | + | <p>Test the results of mating by PCR method. We designed the primers for both <i>MATa</i> and <i>MATα</i> loci. The amplification of both <i>MATa</i> locus and <i>MATα</i> locus indicates that the yeasts has turned into diploids, the <b>MTS</b> has been achieved in other words. </p> |
<hr> | <hr> | ||
| Line 399: | Line 399: | ||
<h4>CONSTRUCTION</h4> | <h4>CONSTRUCTION</h4> | ||
<hr> | <hr> | ||
| − | <p>In the early stage of the project, we constructed two device parts with <i>TEF</i> promoter: BBa_K2407306, BBa_K2407307. At the end of our project, we also constructed one device part with <i>TDH3</i> promoter:BBa_K2407314. Among them, <i>yEmRFP</i> is modified from a mCherry mRFP to adapt to the transcription environment in yeast. We did overlap PCR to | + | <p>In the early stage of the project, we constructed two device parts with <i>TEF</i> promoter: BBa_K2407306, BBa_K2407307. At the end of our project, we also constructed one device part with <i>TDH3</i> promoter:BBa_K2407314. Among them, <i>yEmRFP</i> is modified from a mCherry mRFP to adapt to the transcription environment in yeast. We did overlap PCR to construct them together. After that, we sequenced these parts, and sequencing results showed that these construction were successful.</p> |
<div class="zxx_zoom_demo" align="center"> | <div class="zxx_zoom_demo" align="center"> | ||
<script type="text/javascript" src="https://2017.igem.org/Team:Tianjin/Resources/JS:zoom?action=raw&ctype=text/javascript"></script> | <script type="text/javascript" src="https://2017.igem.org/Team:Tianjin/Resources/JS:zoom?action=raw&ctype=text/javascript"></script> | ||
| Line 405: | Line 405: | ||
<a href="#pic_fortyone"> | <a href="#pic_fortyone"> | ||
<img src="https://static.igem.org/mediawiki/2017/7/71/Tianjin-1-Red_fluorescent_protein_expression_vector_construction_flow_chart.png"></a> | <img src="https://static.igem.org/mediawiki/2017/7/71/Tianjin-1-Red_fluorescent_protein_expression_vector_construction_flow_chart.png"></a> | ||
| − | <p style="font-size:15px;text-align:center"><br/>Fig 2-1. Red fluorescent protein expression vector construction flow chart.</p> | + | <p style="font-size:15px;text-align:center"><br/>Fig.2-1. Red fluorescent protein expression vector construction flow chart.</p> |
</div> | </div> | ||
</div> | </div> | ||
| − | <div id="pic_fortyone" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/b/b9/Tianjin-1-Red_fluorescent_protein_expression_vector_construction_flow_chart_yuan..jpg"><p style="font-size:15px;text-align:center"><br/>Fig 2-1. Red fluorescent protein expression vector construction flow chart.</p></div> | + | <div id="pic_fortyone" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/b/b9/Tianjin-1-Red_fluorescent_protein_expression_vector_construction_flow_chart_yuan..jpg"><p style="font-size:15px;text-align:center"><br/>Fig.2-1. Red fluorescent protein expression vector construction flow chart.</p></div> |
| − | <p>Then we first inserted BBa_K2407306 to the <i>SynⅤ</i> of <i>Saccharomyces cerevisiae</i> . Through the screening of <i>SC-Ura</i> solid medium and PCR experiments, we obtained the required strains called <b><i>PVUVC</i></b>. Second, we integrated the second device part into this chromosome through homologous recombination, allowing the <i>yEmRFP</i> gene to replace the <i>Ura3</i> gene. The <i>5-FOA</i> solid medium and PCR experiments were used to screen correct colony <b><i>PVRVC</i></b>. The insertion of the last | + | <p>Then we first inserted BBa_K2407306 to the <i>SynⅤ</i> of <i>Saccharomyces cerevisiae</i> . Through the screening of <i>SC-Ura</i> solid medium and PCR experiments, we obtained the required strains called <b><i>PVUVC</i></b>. Second, we integrated the second device part into this chromosome through homologous recombination, allowing the <i>yEmRFP</i> gene to replace the <i>Ura3</i> gene. The <i>5-FOA</i> solid medium and PCR experiments were used to screen correct colony <b><i>PVRVC</i></b>. The insertion of the last part referred to the previous method. This process is graphically displayed on the above figure.</p> |
| − | <p>To achieve mating, another mating type of wild haploid yeast <i>Saccharomyces cerevisiae BY4742</i> was used for modification. By digestion and ligation, we | + | <p>To achieve mating, another mating type of wild haploid yeast <i>Saccharomyces cerevisiae BY4742</i> was used for modification. By digestion and ligation, we constructed vika gene on plasmid <i>pRS416</i> which contains a selective marker <i>Ura3</i>, and plasmid <i>pRS413</i> which contains a selective marker <i>His</i>. Then we introduced those two different plasmids into <I>BY4742</I> respectively.</p> |
<h4>Results of Characterization of Mating Switcher</h4> | <h4>Results of Characterization of Mating Switcher</h4> | ||
<hr> | <hr> | ||
<h5>1) Proof of Existence of Device Parts</h5> | <h5>1) Proof of Existence of Device Parts</h5> | ||
| − | <p>We built three device parts in total. They were integrated into the chromosomes of <i>Saccharomyces cerevisiae</i> by transformation. We used colony PCR to | + | <p>We built three device parts in total. They were integrated into the chromosomes of <i>Saccharomyces cerevisiae</i> by transformation. We used colony PCR to prove the existence of these three parts in our strain. The result was showed as below.</p> |
| Line 431: | Line 431: | ||
</div> | </div> | ||
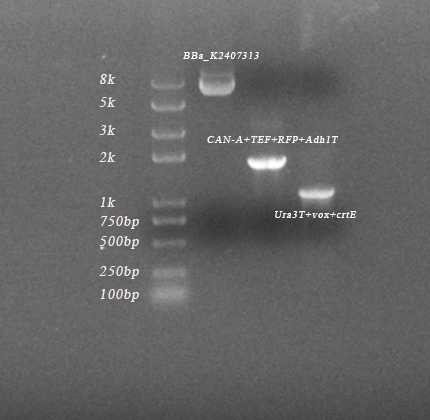
| − | <p style="text-align:center;font-size:1em;">Fig 2-2. | + | <p style="text-align:center;font-size:1em;">Fig.2-2. The results of PCR of <b><i>PVUVC</i></b>, <b><i>TVUVC</i></b>, <b><i>PVUVC</i></b> colonies. (length of 7607bp, 7865bp, 8131bp) As we can see, three parts and all fragments had been amplified, which indicated that we succeeded in constructing them.</p> |
| − | + | ||
| Line 438: | Line 437: | ||
<h5>2) Verification of yEmRFP in the colonies</h5> | <h5>2) Verification of yEmRFP in the colonies</h5> | ||
<hr> | <hr> | ||
| − | <p>The main characterization method of verification of <i>yEmRFP</i> in the <b><i>TVRVC</i></b> applied by us | + | <p>The main characterization method of verification of <i>yEmRFP</i> in the <b><i>TVRVC</i></b> applied by us was observing the expression of <i>red fluorescent protein</i> under the fluorescence microscope. By this way, it will be much more intuitive so that we can directly get the results. We took pictures under different visions and the results are as follows.The experiments of <b><i>PVRVC</i></b> regulation system used this assay method as well.</p> |
<div id="threepic1"> | <div id="threepic1"> | ||
| Line 451: | Line 450: | ||
</div> | </div> | ||
| − | <p style="text-align:center;font-size:1em;">Fig 2-3. Microscope image of yeast | + | <p style="text-align:center;font-size:1em;">Fig.2-3. Microscope image of yeast |
cultured with <i>SC-Leu</i> with <i>yEmRFP</i> gene transformed.</p> | cultured with <i>SC-Leu</i> with <i>yEmRFP</i> gene transformed.</p> | ||
<div id="threepic2"> | <div id="threepic2"> | ||
| Line 461: | Line 460: | ||
</div> | </div> | ||
</div> | </div> | ||
| − | <p style="text-align:center;font-size:1em;">Fig 2-3. Microscope image of yeast | + | <p style="text-align:center;font-size:1em;">Fig.2-3. Microscope image of yeast |
cultured with <i>SC-Leu</i> with <i>yEmRFP</i> gene transformed.</p> | cultured with <i>SC-Leu</i> with <i>yEmRFP</i> gene transformed.</p> | ||
| Line 469: | Line 468: | ||
<h5>3) Result of Mating</h5> | <h5>3) Result of Mating</h5> | ||
<hr> | <hr> | ||
| − | <p>After we got the strain that introduced the <i>yEmRFP</i> gene, we let it mate with another mating type haploid yeast, which had plasmid with <i>vika</i> gene. The result | + | <p>After we got the strain that introduced the <i>yEmRFP</i> gene, we let it mate with another mating type haploid yeast, which had plasmid with <i>vika</i> gene. The result was showed as follows:</p> |
<div class="zxx_zoom_demo_island" align="center"> | <div class="zxx_zoom_demo_island" align="center"> | ||
<script type="text/javascript" src="https://2017.igem.org/Team:Tianjin/Resources/JS:zoom?action=raw&ctype=text/javascript"></script> | <script type="text/javascript" src="https://2017.igem.org/Team:Tianjin/Resources/JS:zoom?action=raw&ctype=text/javascript"></script> | ||
| Line 479: | Line 478: | ||
</div> | </div> | ||
| − | <p style="font-size:15px;text-align:center;margin:0 6em"><br/>Fig 2-4. Three modified colonies and one resulting colony. | + | <p style="font-size:15px;text-align:center;margin:0 6em"><br/>Fig.2-4. Three modified colonies and one resulting colony. |
| − | The upper left corner of the microorganism is synthetic <i>Saccharomyces cerevisiae</i>, we integrated modified fragment into its <i>synthetic chromosome V</i>. (<b><i>PVUVC</i></b>) The upper right corner is also synthetic <i>Saccharomyces cerevisiae</i>. (<b><i>PVRVC</i></b>) It is imported <i>red fluorescent protein</i> gene based on the upper left corner of the yeast. Both of them are single-celled organism called a. The lower right corner of the yeast is another mating type of haploid yeast called α. It has plasmid <i>pRS416</i> with <i>vika</i> gene. The yeast in the lower left corner | + | The upper left corner of the microorganism is synthetic <i>Saccharomyces cerevisiae</i>, we integrated modified fragment into its <i>synthetic chromosome V</i>. (<b><i>PVUVC</i></b>) The upper right corner is also synthetic <i>Saccharomyces cerevisiae</i>. (<b><i>PVRVC</i></b>) It is imported <i>red fluorescent protein</i> gene based on the upper left corner of the yeast. Both of them are single-celled organism called a. The lower right corner of the yeast is another mating type of haploid yeast called α. It has plasmid <i>pRS416</i> with <i>vika</i> gene. The yeast in the lower left corner is diploid <i>Saccharomyces cerevisiae</i>, which is obtained by mating the two yeasts on the right side of the figure.</p> |
| − | <div id="pic_fortythree" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/0/0a/Chenxiyuyuantu2.jpg"><p style="font-size:15px;text-align:center"><br/>Fig 2-4. Three modified colonies and one resulting colony. | + | <div id="pic_fortythree" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/0/0a/Chenxiyuyuantu2.jpg"><p style="font-size:15px;text-align:center"><br/>Fig.2-4. Three modified colonies and one resulting colony. |
| − | <br>The upper left corner of the microorganism is synthetic <i>Saccharomyces cerevisiae</i>, we integrated modified fragment into its <i>synthetic chromosome V</i>. (<b><i>PVUVC</i></b>) The upper right corner is also synthetic <i>Saccharomyces cerevisiae</i>. (<b><i>PVRVC</i></b>) It is imported <i>red fluorescent protein</i> gene based on the upper left corner of the yeast. Both of them are single-celled organism called a. The lower right corner of the yeast is another mating type of haploid yeast called α. It has plasmid <i>pRS416</i> with <i>vika</i> gene. The yeast in the lower left corner | + | <br>The upper left corner of the microorganism is synthetic <i>Saccharomyces cerevisiae</i>, we integrated modified fragment into its <i>synthetic chromosome V</i>. (<b><i>PVUVC</i></b>) The upper right corner is also synthetic <i>Saccharomyces cerevisiae</i>. (<b><i>PVRVC</i></b>) It is imported <i>red fluorescent protein</i> gene based on the upper left corner of the yeast. Both of them are single-celled organism called a. The lower right corner of the yeast is another mating type of haploid yeast called α. It has plasmid <i>pRS416</i> with <i>vika</i> gene. The yeast in the lower left corner is diploid <i>Saccharomyces cerevisiae</i>, which is obtained by mating the two yeasts on the right side of the figure.</br></p></div> |
| − | <p>The yellow colony in the figure | + | <p>The yellow colony in the figure was mating successfully. After the induction of <i>galactose</i>, <i>vika recombinase</i> was expressed, and <i>yEmRFP</i> gene and terminator was deleted so that <i>β-carotene</i> expresses. The color of colony was changed from white to yellow. In addition to it, we also tried other methods to turn on the switch.</p> |
<div class="zxx_zoom_demo_angry" align="center"> | <div class="zxx_zoom_demo_angry" align="center"> | ||
| Line 491: | Line 490: | ||
<a href="#pic_fortyfour"> | <a href="#pic_fortyfour"> | ||
<img src="https://static.igem.org/mediawiki/2017/9/98/Yasuo3333.png"></a> | <img src="https://static.igem.org/mediawiki/2017/9/98/Yasuo3333.png"></a> | ||
| − | <p style="font-size:15px;text-align:center"><br/>Fig 2-5. Yeast after mating cultivated on the Sc-His plate.<br>There are 377 yellow colonies and 365 white colonies in the field of view.</br></p> | + | <p style="font-size:15px;text-align:center"><br/>Fig.2-5. Yeast after mating cultivated on the Sc-His plate.<br>There are 377 yellow colonies and 365 white colonies in the field of view.</br></p> |
</div> | </div> | ||
</div> | </div> | ||
| − | <div id="pic_fortyfour" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/7/77/Tianjin-3-Bacteria_after_mating_cultivated_on_the_Sc-His_plate_yuantu.png"><br/>Fig 2-5. Yeast after mating cultivated on the Sc-His plate. | + | <div id="pic_fortyfour" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/7/77/Tianjin-3-Bacteria_after_mating_cultivated_on_the_Sc-His_plate_yuantu.png"><br/>Fig.2-5. Yeast after mating cultivated on the Sc-His plate. |
<br>There are 377 yellow colonies and 365 white colonies in the field of view.</br></p></div> | <br>There are 377 yellow colonies and 365 white colonies in the field of view.</br></p></div> | ||
| Line 506: | Line 505: | ||
<a href="#pic_fortyfive"> | <a href="#pic_fortyfive"> | ||
<img src="https://static.igem.org/mediawiki/2017/8/81/Tianjin-4-Bacteria_after_mating_cultivated_on_the_Sc-Ura_plate.png"></a> | <img src="https://static.igem.org/mediawiki/2017/8/81/Tianjin-4-Bacteria_after_mating_cultivated_on_the_Sc-Ura_plate.png"></a> | ||
| − | <p style="font-size:15px;text-align:center"><br/>Fig 2-6. | + | <p style="font-size:15px;text-align:center"><br/>Fig.2-6. Yeast after induction cultivated on the Sc-Ura plate. |
<br>There are 325 yellow colonies and 31 white colonies in the field of view.</br></p> | <br>There are 325 yellow colonies and 31 white colonies in the field of view.</br></p> | ||
</div> | </div> | ||
| Line 512: | Line 511: | ||
</div> | </div> | ||
| − | <div id="pic_fortyfive" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/6/6d/Tianjin-4-Bacteria_after_mating_cultivated_on_the_Sc-Ura_plate_yuantu.png"><br/>Fig 2-6. Yeast after induction cultivated on the Sc-Ura plate. | + | <div id="pic_fortyfive" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/6/6d/Tianjin-4-Bacteria_after_mating_cultivated_on_the_Sc-Ura_plate_yuantu.png"><br/>Fig.2-6. Yeast after induction cultivated on the Sc-Ura plate. |
<br>There are 325 yellow colonies and 31 white colonies in the field of view.</br></p></div> | <br>There are 325 yellow colonies and 31 white colonies in the field of view.</br></p></div> | ||
<p>Apart from mating, we also transformed plasmid <i>pRS416</i> with <i>vika</i> gene into the <b><i>PVRVC</i></b>. The efficiency is up to 91.3 percent in this figure.</p> | <p>Apart from mating, we also transformed plasmid <i>pRS416</i> with <i>vika</i> gene into the <b><i>PVRVC</i></b>. The efficiency is up to 91.3 percent in this figure.</p> | ||
| − | <p>Compare above two methods, we | + | <p>Compare above two methods, we found that mating was not as efficient as the transformation of the plasmid. After analysis, we came to the conclusions as follows. For the mating method, <i>vika recombinase</i> has stopped expressing when <i>BY4742</i> mated with <i><b>PVRVC</b></i> in YPD medium. The previously expressed <i>vika recombinase</i> may be degraded during the growth. In contrast to this, with another method that the plasmid was transformed into <i><b>PVRVC</b></i> directly, <i>vika recombinase</i> was continuously expressing during growth. So the efficiency of the second method is higher than the first method.</p> |
<div class="zxx_zoom_demo_qqqqq" align="center"> | <div class="zxx_zoom_demo_qqqqq" align="center"> | ||
| Line 523: | Line 522: | ||
<a href="#pic_fortysix"> | <a href="#pic_fortysix"> | ||
<img src="https://static.igem.org/mediawiki/2017/a/a9/Tianjin-5-Four_modified_colonies_inserted_with_promotor-vox-RFP-terminators-vox-crt_structure.jpg"></a> | <img src="https://static.igem.org/mediawiki/2017/a/a9/Tianjin-5-Four_modified_colonies_inserted_with_promotor-vox-RFP-terminators-vox-crt_structure.jpg"></a> | ||
| − | <p style="font-size:15px;text-align:center"><br/>Fig 2-7. Four modified coloniesinserted with promotor-vox-RFP-terminators-vox-crt structure</p> | + | <p style="font-size:15px;text-align:center"><br/>Fig.2-7. Four modified coloniesinserted with promotor-vox-RFP-terminators-vox-crt structure</p> |
</div> | </div> | ||
</div> | </div> | ||
| − | <div id="pic_fortysix" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/1/1d/Chenxinyuyuantu5.jpg"><br/>Fig 2-7. Four modified coloniesinserted with promotor-vox-RFP-terminators-vox-crt structure</p></div> | + | <div id="pic_fortysix" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/1/1d/Chenxinyuyuantu5.jpg"><br/>Fig.2-7. Four modified coloniesinserted with promotor-vox-RFP-terminators-vox-crt structure</p></div> |
| − | <p>We also used other <i>Saccharomyces cerevisiae</i> with mating type of a to achieve mating switcher. After changing <i>TEF</i> promotor to <i>TDH3</i> promotor, we repeated the test according to the above two methods. The four strains are all haploid synthetic <i>Saccharomyces cerevisiae</i> with mating type of a named <i><b>TVRVC</b></i> NO.2 (upper left), NO.4 (upper right), NO.11 (lower left) and NO.19 (lower right) respectively. The color | + | <p>We also used other <i>Saccharomyces cerevisiae</i> with mating type of a to achieve mating switcher. After changing <i>TEF</i> promotor to <i>TDH3</i> promotor, we repeated the test according to the above two methods. The four strains are all haploid synthetic <i>Saccharomyces cerevisiae</i> with mating type of a named <i><b>TVRVC</b></i> NO.2 (upper left), NO.4 (upper right), NO.11 (lower left) and NO.19 (lower right) respectively. The color appeared to be white because <i>β-carotene</i> did not express.</p> |
<div class="zxx_zoom_demo_qqqqq" align="center"> | <div class="zxx_zoom_demo_qqqqq" align="center"> | ||
| Line 536: | Line 535: | ||
<a href="#pic_fortyseven"> | <a href="#pic_fortyseven"> | ||
<img src="https://static.igem.org/mediawiki/2017/e/e1/Tianjin-6-Four_successful_mating_colonies.jpg"></a> | <img src="https://static.igem.org/mediawiki/2017/e/e1/Tianjin-6-Four_successful_mating_colonies.jpg"></a> | ||
| − | <p style="font-size:15px;text-align:center"><br/>Fig 2-8. Successful mating colonies</p> | + | <p style="font-size:15px;text-align:center"><br/>Fig.2-8. Successful mating colonies</p> |
</div> | </div> | ||
</div> | </div> | ||
| − | <div id="pic_fortyseven" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/1/17/Chenxinyuyuantu6.jpg"><br/>Fig 2-8. Successful mating colonies</p></div> | + | <div id="pic_fortyseven" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/1/17/Chenxinyuyuantu6.jpg"><br/>Fig.2-8. Successful mating colonies</p></div> |
<p>These are parts of successful results of mating mentioned above.</p> | <p>These are parts of successful results of mating mentioned above.</p> | ||
| − | <p>To sum up, the mating switcher can be presented in kinds of yeast with different forms. This | + | <p>To sum up, the mating switcher can be presented in kinds of yeast with different forms. This proved that our Mating switcher was fast, flexible and efficient.</p> |
| − | <p>Meantime, we cultured the transformed yeast in | + | <p>Meantime, we cultured the transformed yeast in 5mL liquid <i>SC-Leu</i> at 30℃ and 220 rpm for 12 hours ( Take three samples at a time). We used one sample for centrifugation to precipitate the yeast and the remaining two remained unchanged. The difference was the fluorescence value we need, then we calculated the average value of them. The excitation wavelength was set at 540nm and the emission wavelength was set at 635nm. Hereafter, we measured the yeast concentration at OD<sub>600</sub>. At last, we divided the fluorescence value by OD<sub>600</sub> to normalize the value and the result data was as follows. |
</p> | </p> | ||
<div class="zxx_zoom_demo_qqqqq" align="center"> | <div class="zxx_zoom_demo_qqqqq" align="center"> | ||
| Line 553: | Line 552: | ||
<a href="#pic_fiftythree"> | <a href="#pic_fiftythree"> | ||
<img src="https://static.igem.org/mediawiki/2017/c/ce/Tianjinzhangshiyu_yasuotu.png"></a> | <img src="https://static.igem.org/mediawiki/2017/c/ce/Tianjinzhangshiyu_yasuotu.png"></a> | ||
| − | <p style="font-size:15px;text-align:center"><br/>Fig 2-9. Normalized fluorescence value was calculated by dividing fluorescent value by cell concentration(OD<sub>600</sub>)</p> | + | <p style="font-size:15px;text-align:center"><br/>Fig.2-9. Normalized fluorescence value was calculated by dividing fluorescent value by cell concentration(OD<sub>600</sub>)</p> |
</div> | </div> | ||
| Line 560: | Line 559: | ||
<div id="pic_fiftythree" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/3/3e/Tianjinzhangshiyu_yuantu.png"><p style="font-size:15px;text-align:center"><br/>Fig.2-9 | <div id="pic_fiftythree" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/3/3e/Tianjinzhangshiyu_yuantu.png"><p style="font-size:15px;text-align:center"><br/>Fig.2-9 | ||
Normalized fluorescence value was calculated by dividing fluorescent value by cell concentration(OD<sub>600</sub>)</p></div> | Normalized fluorescence value was calculated by dividing fluorescent value by cell concentration(OD<sub>600</sub>)</p></div> | ||
| − | <p>From the data we can | + | <p>From the data we can found that the fluorescence intensity of the modified yeast was more than twice that of the wild type. Low red fluorescence was detected after yeast mating, which can be attributed to the influence of <i>β-carotene</i>.</p> |
<h4>DISCUSSION & FUTURE WORK</h4> | <h4>DISCUSSION & FUTURE WORK</h4> | ||
<hr> | <hr> | ||
| − | <p>Our mating switch plays an important role in many respects, such as including heavy metal treatment and cell signal switching. And we created a novel method to prove the effectiveness of the switch in an intuitive and effective way. The terminator of the first part (PVUVC) terminates the expression of the downstream gene, proving the validity of the switcher, and the second part (<b><i>PVRVC</i></b>) creates an evident method of color conversion to determine the state of the switcher.</p> | + | <p>Our mating switch plays an important role in many respects, such as including heavy metal treatment and cell signal switching. And we created a novel method to prove the effectiveness of the switch in an intuitive and effective way. The terminator of the first part (<b><i>PVUVC</i></b>) terminates the expression of the downstream gene, proving the validity of the switcher, and the second part (<b><i>PVRVC</i></b>) creates an evident method of color conversion to determine the state of the switcher.</p> |
| − | <p>Aiming to increase the Vika-vox system efficiency, we let Vika enzyme saturate expression, but the efficiency was still relatively low. We hypothesized that this phenomenon was caused by degradation of the Vika enzyme in the YPD culture medium. We’d better change the composition or proportion of YPD ingredients to find out the best culture conditions. We are looking forward to more research in this field so that we can make this system work better and even perfectly.</p> | + | <p>Aiming to increase the <i>Vika-vox</i> system efficiency, we let <i>Vika</i> enzyme saturate expression, but the efficiency was still relatively low. We hypothesized that this phenomenon was caused by degradation of the <i>Vika</i> enzyme in the YPD culture medium. We’d better change the composition or proportion of YPD ingredients to find out the best culture conditions. We are looking forward to more research in this field so that we can make this system work better and even perfectly.</p> |
<p>We use the <i>RFP</i> as the reporting protein. But there exists a drawback that it’s detected with an expensive device. A more intuitive reporting strategy need to be developed, maybe it can be seen by bare eyes like <i>E.coli</i> in the near future.</p> | <p>We use the <i>RFP</i> as the reporting protein. But there exists a drawback that it’s detected with an expensive device. A more intuitive reporting strategy need to be developed, maybe it can be seen by bare eyes like <i>E.coli</i> in the near future.</p> | ||
| Line 1,077: | Line 1,076: | ||
<a href="#pic_seventy-five"> | <a href="#pic_seventy-five"> | ||
<img src="https://static.igem.org/mediawiki/2017/b/b7/Heavy-metal-jiaotu.jpg"></a> | <img src="https://static.igem.org/mediawiki/2017/b/b7/Heavy-metal-jiaotu.jpg"></a> | ||
| − | <p style="font-size:15px;text-align:center"><br/> | + | <p style="font-size:15px;text-align:center"><br/>Fig.5-1 The results of PCR of our <i>S.C-Cu</i>. <i>LIMT</i> gene (length of 319bp) 、<i>Cup1</i>(length of 186bp) and complete sequence(length of 3114bp)have been amplified. which indicated that we succeeded in the construction of genetic circuit.</p> |
</div> | </div> | ||
</div> | </div> | ||
| − | <div id="pic_seventy-five" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/b/b7/Heavy-metal-jiaotu.jpg"><p style="font-size:15px;text-align:center"><br/> | + | <div id="pic_seventy-five" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/b/b7/Heavy-metal-jiaotu.jpg"><p style="font-size:15px;text-align:center"><br/>Fig.5-1 The results of PCR of our <i>S.C-Cu</i>. <i>LIMT</i> gene (length of 319bp) 、<i>Cup1</i>(length of 186bp) and complete sequence(length of 3114bp)have been amplified. which indicated that we succeeded in the construction of genetic circuit.</p></div> |
<p>Fig.5-1 the results of PCR. We use <i>2k plus Ⅱ</i> as the marker. On four parallel lanes of the gel (number 1,2,3,4), run were four set of DNA molecules of known size (319bp for number 1, the <i>LIMT</i>; 186bp for number 2 and 3, the <i>Cup1</i>; 3114bp for number 4,the whole sequence contained <i>Cup1</i>). From the DNA band of number 1, we could analyze that <i>vika</i> has been expressed to delete the <i>Cup1</i> and its terminor, so we can get the <i>LIMT</i>. From the DNA band of number 2, 3 and 4, we could delightedly prove that the fragments (<i>TEF</i> promoter, <i>Cup1</i> and <i>ura3</i> terminator) have successfully transformed to synthetic chromosome <i>V</i>. </p> | <p>Fig.5-1 the results of PCR. We use <i>2k plus Ⅱ</i> as the marker. On four parallel lanes of the gel (number 1,2,3,4), run were four set of DNA molecules of known size (319bp for number 1, the <i>LIMT</i>; 186bp for number 2 and 3, the <i>Cup1</i>; 3114bp for number 4,the whole sequence contained <i>Cup1</i>). From the DNA band of number 1, we could analyze that <i>vika</i> has been expressed to delete the <i>Cup1</i> and its terminor, so we can get the <i>LIMT</i>. From the DNA band of number 2, 3 and 4, we could delightedly prove that the fragments (<i>TEF</i> promoter, <i>Cup1</i> and <i>ura3</i> terminator) have successfully transformed to synthetic chromosome <i>V</i>. </p> | ||
| Line 1,096: | Line 1,095: | ||
<div class="small_pic_demo_qqqqqqq" style="float:left;"> | <div class="small_pic_demo_qqqqqqq" style="float:left;"> | ||
<a href="#pic_Seventy-six"> | <a href="#pic_Seventy-six"> | ||
| − | <img src="https://static.igem.org/mediawiki/parts/1/17/Demonstrate.Cu.png"></a><p style="font-size:15px;text-align:center"><br/> | + | <img src="https://static.igem.org/mediawiki/parts/1/17/Demonstrate.Cu.png"></a><p style="font-size:15px;text-align:center"><br/>Fig.5-2 The variations of copper(II) consumption with time for <i>S.C-Cu</i>、<i>S8</i> and <i>BY4741</i> at 430 mg/L copper(II) concentrations. |
</p> | </p> | ||
</div> | </div> | ||
| Line 1,102: | Line 1,101: | ||
<a href="#pic_Seventy-seven" > | <a href="#pic_Seventy-seven" > | ||
<img src="https://static.igem.org/mediawiki/parts/5/5f/Demonstrate.Cd.png"/> | <img src="https://static.igem.org/mediawiki/parts/5/5f/Demonstrate.Cd.png"/> | ||
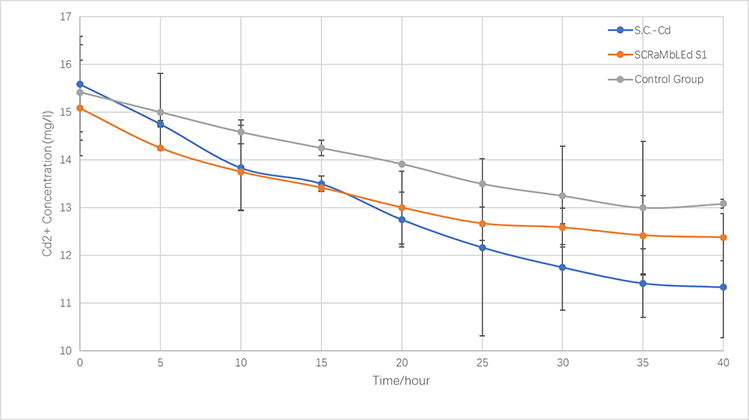
| − | </a> <p style="font-size:15px;text-align:center"><br/> | + | </a> <p style="font-size:15px;text-align:center"><br/>Fig.5-3 The variations of cadmium(II) consumption with time for <i>S.C-Cd</i>、<i>S1</i> and <i>BY4741</i> at 16 mg/L cadmium(II) concentrations</p> |
</div> | </div> | ||
</div> | </div> | ||
| − | <div id="pic_Seventy-six" style="display:none;"><img src="https://static.igem.org/mediawiki/parts/1/17/Demonstrate.Cu.png"/><p style="font-size:15px;text-align:center"><br/>The variations of copper(II) consumption with time for <i>S.C-Cu</i>、<i>S8</i> and <i>BY4741</i> at 430 mg/L copper(II) concentrations. | + | <div id="pic_Seventy-six" style="display:none;"><img src="https://static.igem.org/mediawiki/parts/1/17/Demonstrate.Cu.png"/><p style="font-size:15px;text-align:center"><br/>Fig.5-2 The variations of copper(II) consumption with time for <i>S.C-Cu</i>、<i>S8</i> and <i>BY4741</i> at 430 mg/L copper(II) concentrations. |
</p></div> | </p></div> | ||
| − | <div id="pic_Seventy-seven" style="display:none;"><img src="https://static.igem.org/mediawiki/parts/5/5f/Demonstrate.Cd.png"/><p style="font-size:15px;text-align:center"><br/> | + | <div id="pic_Seventy-seven" style="display:none;"><img src="https://static.igem.org/mediawiki/parts/5/5f/Demonstrate.Cd.png"/><p style="font-size:15px;text-align:center"><br/>Fig.5-3 The variations of cadmium(II) consumption with time for <i>S.C-Cd</i>、<i>S1</i> and <i>BY4741</i> at 16 mg/L cadmium(II) concentrations </p></div> |
<p>Afterwards, we check if the <i>vika</i> enzyme could work well. The Cu yeast with a plasmid expressing <i>vika</i> enzyme is grew in the medium with <i> raffinose</i>, then transferred to heavy metal solution. | <p>Afterwards, we check if the <i>vika</i> enzyme could work well. The Cu yeast with a plasmid expressing <i>vika</i> enzyme is grew in the medium with <i> raffinose</i>, then transferred to heavy metal solution. | ||
<div class="zxx_zoom_demo_qqqqqqq" align="center"> | <div class="zxx_zoom_demo_qqqqqqq" align="center"> | ||
| Line 1,115: | Line 1,114: | ||
<a href="#pic_eighty"> | <a href="#pic_eighty"> | ||
<img src="https://static.igem.org/mediawiki/2017/6/6b/Design.cu-cd.curve.png"></a> | <img src="https://static.igem.org/mediawiki/2017/6/6b/Design.cu-cd.curve.png"></a> | ||
| − | <p style="font-size:15px;text-align:center"><br/> | + | <p style="font-size:15px;text-align:center"><br/>Fig.5-4 <i>S.C-Cu</i> is cultivated in medium with <i> raffinose </i> including 320mg/L copper ions and 6mg/L Cadmium.<i>galactose</i> is added at 12 hours to turn on "switch".</p> |
</div> | </div> | ||
</div> | </div> | ||
| − | <div id="pic_eighty" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/6/6b/Design.cu-cd.curve.png"><p style="font-size:15px;text-align:center"><br/> | + | <div id="pic_eighty" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/6/6b/Design.cu-cd.curve.png"><p style="font-size:15px;text-align:center"><br/>Fig.5-4 <i>S.C-Cu</i> is cultivated in medium with <i> raffinose </i> including 320mg/L copper ions and 6mg/L Cadmium.<i>galactose</i> is added at 12 hours to turn on "switch".</p></div> |
<p>Fig.5-4 clearly shows the change of the concentration of heavy metal ions in the supernatant. Firstly, the Cu yeast works smoothly. The concentration of copper ions declines over time while that of cadmium ions barely changes. 12 hours later, we add <i>galactose</i> to the solution. Situation changes. <i>Galactose</i> induces the enzyme, changing Cu yeast to Cd yeast. It leads to faster adsorption of cadmium but slower for copper.</p> | <p>Fig.5-4 clearly shows the change of the concentration of heavy metal ions in the supernatant. Firstly, the Cu yeast works smoothly. The concentration of copper ions declines over time while that of cadmium ions barely changes. 12 hours later, we add <i>galactose</i> to the solution. Situation changes. <i>Galactose</i> induces the enzyme, changing Cu yeast to Cd yeast. It leads to faster adsorption of cadmium but slower for copper.</p> | ||
| Line 1,138: | Line 1,137: | ||
<div class="collapse-card__close_handler mt1 align-right mouse-pointer"> | <div class="collapse-card__close_handler mt1 align-right mouse-pointer"> | ||
| − | + | go bottom and find a surprise <i class="fa fa-chevron-up"></i> | |
</div> | </div> | ||
</div> | </div> | ||
Latest revision as of 03:27, 2 November 2017
/* OVERRIDE IGEM SETTINGS */