Zsy5691750 (Talk | contribs) |
Cccxyyyyyyyy (Talk | contribs) |
||
| (7 intermediate revisions by 3 users not shown) | |||
| Line 254: | Line 254: | ||
<div id="title"> | <div id="title"> | ||
<h1>Demonstrate</h1> | <h1>Demonstrate</h1> | ||
| − | <hr> | + | <a title="huge suprise" href="https://2017.igem.org/Team:Tianjin/surprise23333" target="_blank"><hr></a> |
</div> | </div> | ||
| Line 282: | Line 282: | ||
<a href="#pic_eleven" > | <a href="#pic_eleven" > | ||
<img src="https://static.igem.org/mediawiki/2017/0/03/Tianjin-ho-result-9.jpeg"/> | <img src="https://static.igem.org/mediawiki/2017/0/03/Tianjin-ho-result-9.jpeg"/> | ||
| − | </a> <p style="font-size:15px;text-align:center"><br/>Fig. 1-2. As we can see in the gel photo above, the <b>UP</b> and <b>DOWN</b> segments hasn’t been amplified in our <i><b>SynⅩ-dUra</b></i> comparing to the BY4741 as control | + | </a> <p style="font-size:15px;text-align:center"><br/>Fig. 1-2. As we can see in the gel photo above, the <b>UP</b> and <b>DOWN</b> segments hasn’t been amplified in our <i><b>SynⅩ-dUra</b></i> comparing to the BY4741 as control, which indicated that the HMRa gene has been successfully eliminated. |
</p> | </p> | ||
</div> | </div> | ||
| Line 289: | Line 289: | ||
<div id="pic_ten" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/1/1b/Tianjin-ho-result-10.jpeg"/><p style="font-size:15px;text-align:center"><br/> Fig. 1-1. The PCR strategy for testing whether we deleted the HMRa in <b><i>SynⅩ-dUra</i></b>. | <div id="pic_ten" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/1/1b/Tianjin-ho-result-10.jpeg"/><p style="font-size:15px;text-align:center"><br/> Fig. 1-1. The PCR strategy for testing whether we deleted the HMRa in <b><i>SynⅩ-dUra</i></b>. | ||
</p></div> | </p></div> | ||
| − | <div id="pic_eleven" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/0/03/Tianjin-ho-result-9.jpeg"/><p style="font-size:15px;text-align:center"><br/>Fig. 1-2. As we can see in the gel photo above, the <b>UP</b> and <b>DOWN</b> segments hasn’t been amplified in our <i><b>SynⅩ-dUra</b></i> comparing to the BY4741 as control | + | <div id="pic_eleven" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/0/03/Tianjin-ho-result-9.jpeg"/><p style="font-size:15px;text-align:center"><br/>Fig. 1-2. As we can see in the gel photo above, the <b>UP</b> and <b>DOWN</b> segments hasn’t been amplified in our <i><b>SynⅩ-dUra</b></i> comparing to the BY4741 as control, which indicated that the HMRa gene has been successfully eliminated. |
</p></div> | </p></div> | ||
<h4> The result for constructing the <i>Gal</i> systems</h4> | <h4> The result for constructing the <i>Gal</i> systems</h4> | ||
| Line 328: | Line 328: | ||
<p> Cultivate two groups of yeasts together. (one is <b><i>SynⅩ-dUra-416</i></b>, the other is normal <i>BY4741 MATa</i>) If the <b>MTS</b> has been accomplished (<b><i>SynⅩ-dUra-416</i></b> can become <i>MATα</i>), the two groups of haploids can mate with each other and become diploids. </p> | <p> Cultivate two groups of yeasts together. (one is <b><i>SynⅩ-dUra-416</i></b>, the other is normal <i>BY4741 MATa</i>) If the <b>MTS</b> has been accomplished (<b><i>SynⅩ-dUra-416</i></b> can become <i>MATα</i>), the two groups of haploids can mate with each other and become diploids. </p> | ||
<h5>3) Step three</h5> | <h5>3) Step three</h5> | ||
| − | <p>Test the results of mating by PCR method. We designed the primers for both <i>MATa</i> | + | <p>Test the results of mating by PCR method. We designed the primers for both <i>MATa</i> and <i>MATα</i> loci. The amplification of both <i>MATa</i> locus and <i>MATα</i> locus indicates that the yeasts has turned into diploids, the <b>MTS</b> has been achieved in other words. </p> |
<hr> | <hr> | ||
| Line 405: | Line 405: | ||
<a href="#pic_fortyone"> | <a href="#pic_fortyone"> | ||
<img src="https://static.igem.org/mediawiki/2017/7/71/Tianjin-1-Red_fluorescent_protein_expression_vector_construction_flow_chart.png"></a> | <img src="https://static.igem.org/mediawiki/2017/7/71/Tianjin-1-Red_fluorescent_protein_expression_vector_construction_flow_chart.png"></a> | ||
| − | <p style="font-size:15px;text-align:center"><br/>Fig 2-1. Red fluorescent protein expression vector construction flow chart.</p> | + | <p style="font-size:15px;text-align:center"><br/>Fig.2-1. Red fluorescent protein expression vector construction flow chart.</p> |
</div> | </div> | ||
</div> | </div> | ||
| − | <div id="pic_fortyone" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/b/b9/Tianjin-1-Red_fluorescent_protein_expression_vector_construction_flow_chart_yuan..jpg"><p style="font-size:15px;text-align:center"><br/>Fig 2-1. Red fluorescent protein expression vector construction flow chart.</p></div> | + | <div id="pic_fortyone" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/b/b9/Tianjin-1-Red_fluorescent_protein_expression_vector_construction_flow_chart_yuan..jpg"><p style="font-size:15px;text-align:center"><br/>Fig.2-1. Red fluorescent protein expression vector construction flow chart.</p></div> |
<p>Then we first inserted BBa_K2407306 to the <i>SynⅤ</i> of <i>Saccharomyces cerevisiae</i> . Through the screening of <i>SC-Ura</i> solid medium and PCR experiments, we obtained the required strains called <b><i>PVUVC</i></b>. Second, we integrated the second device part into this chromosome through homologous recombination, allowing the <i>yEmRFP</i> gene to replace the <i>Ura3</i> gene. The <i>5-FOA</i> solid medium and PCR experiments were used to screen correct colony <b><i>PVRVC</i></b>. The insertion of the last part referred to the previous method. This process is graphically displayed on the above figure.</p> | <p>Then we first inserted BBa_K2407306 to the <i>SynⅤ</i> of <i>Saccharomyces cerevisiae</i> . Through the screening of <i>SC-Ura</i> solid medium and PCR experiments, we obtained the required strains called <b><i>PVUVC</i></b>. Second, we integrated the second device part into this chromosome through homologous recombination, allowing the <i>yEmRFP</i> gene to replace the <i>Ura3</i> gene. The <i>5-FOA</i> solid medium and PCR experiments were used to screen correct colony <b><i>PVRVC</i></b>. The insertion of the last part referred to the previous method. This process is graphically displayed on the above figure.</p> | ||
| Line 431: | Line 431: | ||
</div> | </div> | ||
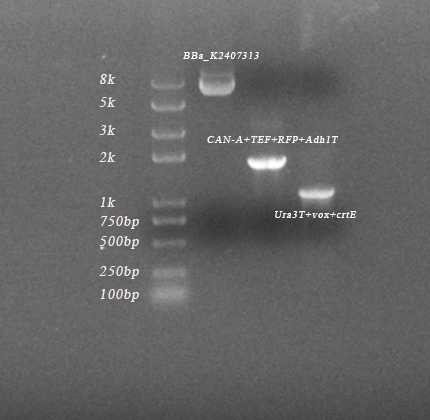
| − | <p style="text-align:center;font-size:1em;">Fig 2-2. | + | <p style="text-align:center;font-size:1em;">Fig.2-2. The results of PCR of <b><i>PVUVC</i></b>, <b><i>TVUVC</i></b>, <b><i>PVUVC</i></b> colonies. (length of 7607bp, 7865bp, 8131bp) As we can see, three parts and all fragments had been amplified, which indicated that we succeeded in constructing them.</p> |
| − | + | ||
| Line 451: | Line 450: | ||
</div> | </div> | ||
| − | <p style="text-align:center;font-size:1em;">Fig 2-3. Microscope image of yeast | + | <p style="text-align:center;font-size:1em;">Fig.2-3. Microscope image of yeast |
cultured with <i>SC-Leu</i> with <i>yEmRFP</i> gene transformed.</p> | cultured with <i>SC-Leu</i> with <i>yEmRFP</i> gene transformed.</p> | ||
<div id="threepic2"> | <div id="threepic2"> | ||
| Line 461: | Line 460: | ||
</div> | </div> | ||
</div> | </div> | ||
| − | <p style="text-align:center;font-size:1em;">Fig 2-3. Microscope image of yeast | + | <p style="text-align:center;font-size:1em;">Fig.2-3. Microscope image of yeast |
cultured with <i>SC-Leu</i> with <i>yEmRFP</i> gene transformed.</p> | cultured with <i>SC-Leu</i> with <i>yEmRFP</i> gene transformed.</p> | ||
| Line 479: | Line 478: | ||
</div> | </div> | ||
| − | <p style="font-size:15px;text-align:center;margin:0 6em"><br/>Fig 2-4. Three modified colonies and one resulting colony. | + | <p style="font-size:15px;text-align:center;margin:0 6em"><br/>Fig.2-4. Three modified colonies and one resulting colony. |
The upper left corner of the microorganism is synthetic <i>Saccharomyces cerevisiae</i>, we integrated modified fragment into its <i>synthetic chromosome V</i>. (<b><i>PVUVC</i></b>) The upper right corner is also synthetic <i>Saccharomyces cerevisiae</i>. (<b><i>PVRVC</i></b>) It is imported <i>red fluorescent protein</i> gene based on the upper left corner of the yeast. Both of them are single-celled organism called a. The lower right corner of the yeast is another mating type of haploid yeast called α. It has plasmid <i>pRS416</i> with <i>vika</i> gene. The yeast in the lower left corner is diploid <i>Saccharomyces cerevisiae</i>, which is obtained by mating the two yeasts on the right side of the figure.</p> | The upper left corner of the microorganism is synthetic <i>Saccharomyces cerevisiae</i>, we integrated modified fragment into its <i>synthetic chromosome V</i>. (<b><i>PVUVC</i></b>) The upper right corner is also synthetic <i>Saccharomyces cerevisiae</i>. (<b><i>PVRVC</i></b>) It is imported <i>red fluorescent protein</i> gene based on the upper left corner of the yeast. Both of them are single-celled organism called a. The lower right corner of the yeast is another mating type of haploid yeast called α. It has plasmid <i>pRS416</i> with <i>vika</i> gene. The yeast in the lower left corner is diploid <i>Saccharomyces cerevisiae</i>, which is obtained by mating the two yeasts on the right side of the figure.</p> | ||
| − | <div id="pic_fortythree" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/0/0a/Chenxiyuyuantu2.jpg"><p style="font-size:15px;text-align:center"><br/>Fig 2-4. Three modified colonies and one resulting colony. | + | <div id="pic_fortythree" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/0/0a/Chenxiyuyuantu2.jpg"><p style="font-size:15px;text-align:center"><br/>Fig.2-4. Three modified colonies and one resulting colony. |
<br>The upper left corner of the microorganism is synthetic <i>Saccharomyces cerevisiae</i>, we integrated modified fragment into its <i>synthetic chromosome V</i>. (<b><i>PVUVC</i></b>) The upper right corner is also synthetic <i>Saccharomyces cerevisiae</i>. (<b><i>PVRVC</i></b>) It is imported <i>red fluorescent protein</i> gene based on the upper left corner of the yeast. Both of them are single-celled organism called a. The lower right corner of the yeast is another mating type of haploid yeast called α. It has plasmid <i>pRS416</i> with <i>vika</i> gene. The yeast in the lower left corner is diploid <i>Saccharomyces cerevisiae</i>, which is obtained by mating the two yeasts on the right side of the figure.</br></p></div> | <br>The upper left corner of the microorganism is synthetic <i>Saccharomyces cerevisiae</i>, we integrated modified fragment into its <i>synthetic chromosome V</i>. (<b><i>PVUVC</i></b>) The upper right corner is also synthetic <i>Saccharomyces cerevisiae</i>. (<b><i>PVRVC</i></b>) It is imported <i>red fluorescent protein</i> gene based on the upper left corner of the yeast. Both of them are single-celled organism called a. The lower right corner of the yeast is another mating type of haploid yeast called α. It has plasmid <i>pRS416</i> with <i>vika</i> gene. The yeast in the lower left corner is diploid <i>Saccharomyces cerevisiae</i>, which is obtained by mating the two yeasts on the right side of the figure.</br></p></div> | ||
| Line 491: | Line 490: | ||
<a href="#pic_fortyfour"> | <a href="#pic_fortyfour"> | ||
<img src="https://static.igem.org/mediawiki/2017/9/98/Yasuo3333.png"></a> | <img src="https://static.igem.org/mediawiki/2017/9/98/Yasuo3333.png"></a> | ||
| − | <p style="font-size:15px;text-align:center"><br/>Fig 2-5. Yeast after mating cultivated on the Sc-His plate.<br>There are 377 yellow colonies and 365 white colonies in the field of view.</br></p> | + | <p style="font-size:15px;text-align:center"><br/>Fig.2-5. Yeast after mating cultivated on the Sc-His plate.<br>There are 377 yellow colonies and 365 white colonies in the field of view.</br></p> |
</div> | </div> | ||
</div> | </div> | ||
| − | <div id="pic_fortyfour" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/7/77/Tianjin-3-Bacteria_after_mating_cultivated_on_the_Sc-His_plate_yuantu.png"><br/>Fig 2-5. Yeast after mating cultivated on the Sc-His plate. | + | <div id="pic_fortyfour" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/7/77/Tianjin-3-Bacteria_after_mating_cultivated_on_the_Sc-His_plate_yuantu.png"><br/>Fig.2-5. Yeast after mating cultivated on the Sc-His plate. |
<br>There are 377 yellow colonies and 365 white colonies in the field of view.</br></p></div> | <br>There are 377 yellow colonies and 365 white colonies in the field of view.</br></p></div> | ||
| Line 506: | Line 505: | ||
<a href="#pic_fortyfive"> | <a href="#pic_fortyfive"> | ||
<img src="https://static.igem.org/mediawiki/2017/8/81/Tianjin-4-Bacteria_after_mating_cultivated_on_the_Sc-Ura_plate.png"></a> | <img src="https://static.igem.org/mediawiki/2017/8/81/Tianjin-4-Bacteria_after_mating_cultivated_on_the_Sc-Ura_plate.png"></a> | ||
| − | <p style="font-size:15px;text-align:center"><br/>Fig 2-6. | + | <p style="font-size:15px;text-align:center"><br/>Fig.2-6. Yeast after induction cultivated on the Sc-Ura plate. |
<br>There are 325 yellow colonies and 31 white colonies in the field of view.</br></p> | <br>There are 325 yellow colonies and 31 white colonies in the field of view.</br></p> | ||
</div> | </div> | ||
| Line 512: | Line 511: | ||
</div> | </div> | ||
| − | <div id="pic_fortyfive" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/6/6d/Tianjin-4-Bacteria_after_mating_cultivated_on_the_Sc-Ura_plate_yuantu.png"><br/>Fig 2-6. Yeast after induction cultivated on the Sc-Ura plate. | + | <div id="pic_fortyfive" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/6/6d/Tianjin-4-Bacteria_after_mating_cultivated_on_the_Sc-Ura_plate_yuantu.png"><br/>Fig.2-6. Yeast after induction cultivated on the Sc-Ura plate. |
<br>There are 325 yellow colonies and 31 white colonies in the field of view.</br></p></div> | <br>There are 325 yellow colonies and 31 white colonies in the field of view.</br></p></div> | ||
| Line 523: | Line 522: | ||
<a href="#pic_fortysix"> | <a href="#pic_fortysix"> | ||
<img src="https://static.igem.org/mediawiki/2017/a/a9/Tianjin-5-Four_modified_colonies_inserted_with_promotor-vox-RFP-terminators-vox-crt_structure.jpg"></a> | <img src="https://static.igem.org/mediawiki/2017/a/a9/Tianjin-5-Four_modified_colonies_inserted_with_promotor-vox-RFP-terminators-vox-crt_structure.jpg"></a> | ||
| − | <p style="font-size:15px;text-align:center"><br/>Fig 2-7. Four modified coloniesinserted with promotor-vox-RFP-terminators-vox-crt structure</p> | + | <p style="font-size:15px;text-align:center"><br/>Fig.2-7. Four modified coloniesinserted with promotor-vox-RFP-terminators-vox-crt structure</p> |
</div> | </div> | ||
</div> | </div> | ||
| − | <div id="pic_fortysix" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/1/1d/Chenxinyuyuantu5.jpg"><br/>Fig 2-7. Four modified coloniesinserted with promotor-vox-RFP-terminators-vox-crt structure</p></div> | + | <div id="pic_fortysix" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/1/1d/Chenxinyuyuantu5.jpg"><br/>Fig.2-7. Four modified coloniesinserted with promotor-vox-RFP-terminators-vox-crt structure</p></div> |
<p>We also used other <i>Saccharomyces cerevisiae</i> with mating type of a to achieve mating switcher. After changing <i>TEF</i> promotor to <i>TDH3</i> promotor, we repeated the test according to the above two methods. The four strains are all haploid synthetic <i>Saccharomyces cerevisiae</i> with mating type of a named <i><b>TVRVC</b></i> NO.2 (upper left), NO.4 (upper right), NO.11 (lower left) and NO.19 (lower right) respectively. The color appeared to be white because <i>β-carotene</i> did not express.</p> | <p>We also used other <i>Saccharomyces cerevisiae</i> with mating type of a to achieve mating switcher. After changing <i>TEF</i> promotor to <i>TDH3</i> promotor, we repeated the test according to the above two methods. The four strains are all haploid synthetic <i>Saccharomyces cerevisiae</i> with mating type of a named <i><b>TVRVC</b></i> NO.2 (upper left), NO.4 (upper right), NO.11 (lower left) and NO.19 (lower right) respectively. The color appeared to be white because <i>β-carotene</i> did not express.</p> | ||
| Line 536: | Line 535: | ||
<a href="#pic_fortyseven"> | <a href="#pic_fortyseven"> | ||
<img src="https://static.igem.org/mediawiki/2017/e/e1/Tianjin-6-Four_successful_mating_colonies.jpg"></a> | <img src="https://static.igem.org/mediawiki/2017/e/e1/Tianjin-6-Four_successful_mating_colonies.jpg"></a> | ||
| − | <p style="font-size:15px;text-align:center"><br/>Fig 2-8. Successful mating colonies</p> | + | <p style="font-size:15px;text-align:center"><br/>Fig.2-8. Successful mating colonies</p> |
</div> | </div> | ||
</div> | </div> | ||
| − | <div id="pic_fortyseven" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/1/17/Chenxinyuyuantu6.jpg"><br/>Fig 2-8. Successful mating colonies</p></div> | + | <div id="pic_fortyseven" style="display:none;"><img src="https://static.igem.org/mediawiki/2017/1/17/Chenxinyuyuantu6.jpg"><br/>Fig.2-8. Successful mating colonies</p></div> |
<p>These are parts of successful results of mating mentioned above.</p> | <p>These are parts of successful results of mating mentioned above.</p> | ||
| Line 553: | Line 552: | ||
<a href="#pic_fiftythree"> | <a href="#pic_fiftythree"> | ||
<img src="https://static.igem.org/mediawiki/2017/c/ce/Tianjinzhangshiyu_yasuotu.png"></a> | <img src="https://static.igem.org/mediawiki/2017/c/ce/Tianjinzhangshiyu_yasuotu.png"></a> | ||
| − | <p style="font-size:15px;text-align:center"><br/>Fig 2-9. Normalized fluorescence value was calculated by dividing fluorescent value by cell concentration(OD<sub>600</sub>)</p> | + | <p style="font-size:15px;text-align:center"><br/>Fig.2-9. Normalized fluorescence value was calculated by dividing fluorescent value by cell concentration(OD<sub>600</sub>)</p> |
</div> | </div> | ||
| Line 1,138: | Line 1,137: | ||
<div class="collapse-card__close_handler mt1 align-right mouse-pointer"> | <div class="collapse-card__close_handler mt1 align-right mouse-pointer"> | ||
| − | + | go bottom and find a surprise <i class="fa fa-chevron-up"></i> | |
</div> | </div> | ||
</div> | </div> | ||
Latest revision as of 03:27, 2 November 2017
/* OVERRIDE IGEM SETTINGS */