| Line 25: | Line 25: | ||
Microfluidics were divide into three parts: the selective chip; the Gaussian chip; and the immobilization chip. | Microfluidics were divide into three parts: the selective chip; the Gaussian chip; and the immobilization chip. | ||
| − | == | + | == The Selection of the <i>Caenorhabditis elegans</i> == |
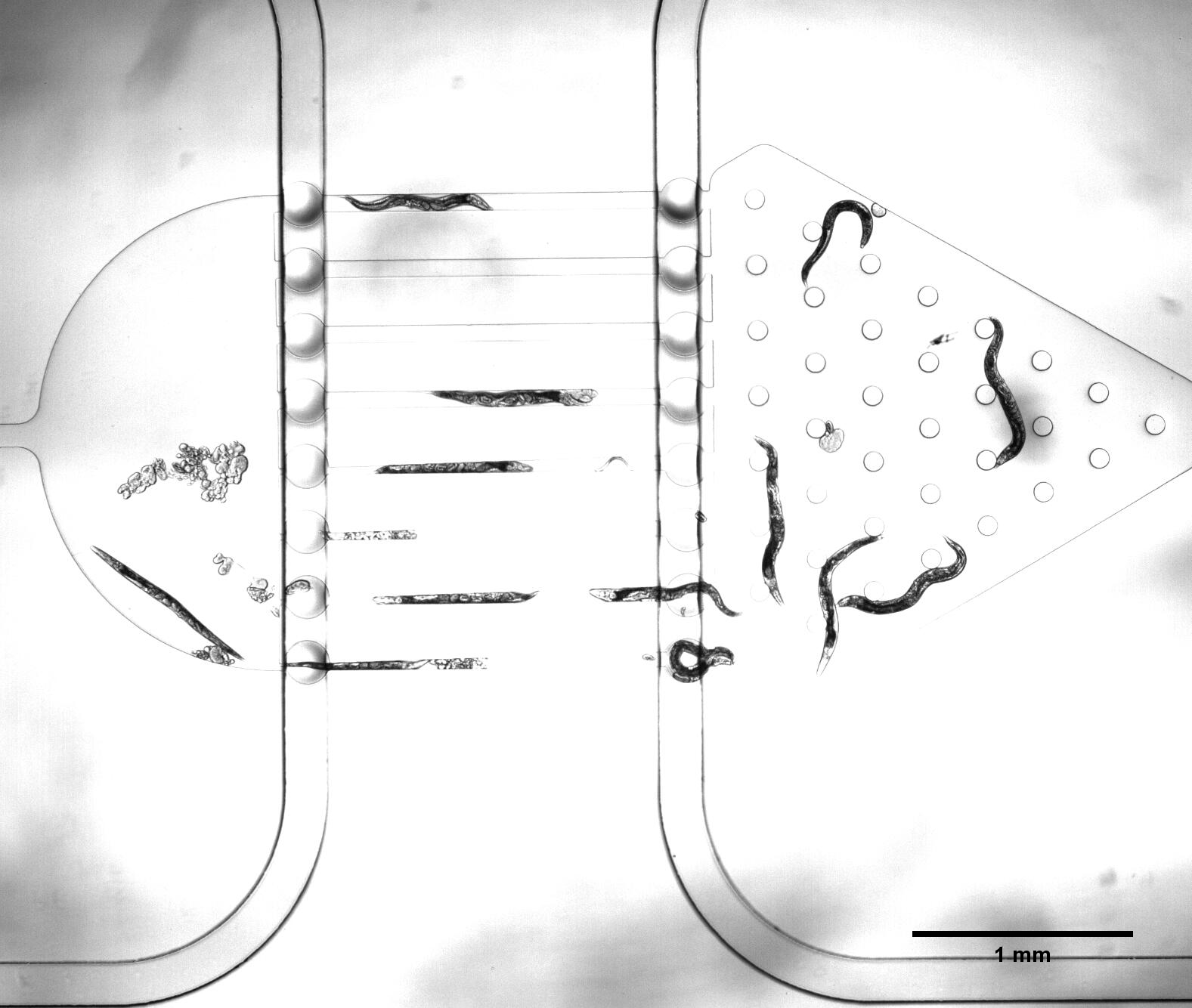
| − | The selective chip | + | There are two plans of selecting worms. <html><a href="https://2017.igem.org/Team:SUSTech_Shenzhen/Hardware">Why we need the C. elegans with the same stage</a></html> The first one is using microfluidics. We designed the selective chip to select the <i>Caenorhabditis elegans (C. elegans)</i> with the appropriate size (Fig.1). |
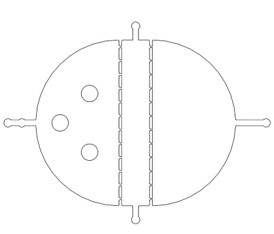
{{SUSTech_Image_Center_8 | filename=T--SUSTech_Shenzhen--Microfuildics--selective.jpeg|width=600px|caption=<B>Fig.1 The Selective Chip</B>}} | {{SUSTech_Image_Center_8 | filename=T--SUSTech_Shenzhen--Microfuildics--selective.jpeg|width=600px|caption=<B>Fig.1 The Selective Chip</B>}} | ||
| − | We | + | We need a large number of the worms with the same stage to do the Gaussion chip. <html><a href="https://2017.igem.org/Team:SUSTech_Shenzhen/Model3">Why we need so many with the same stage</a></html> However, we found that the chip only has 12 fences (Fig.2) and the efficiency of the selective chip was very low because of such a small number of the fences. In addition, the <i>C. elegans</i> have flexible body, some of the suitable size worms would still go through the second fences (Fig.2). |
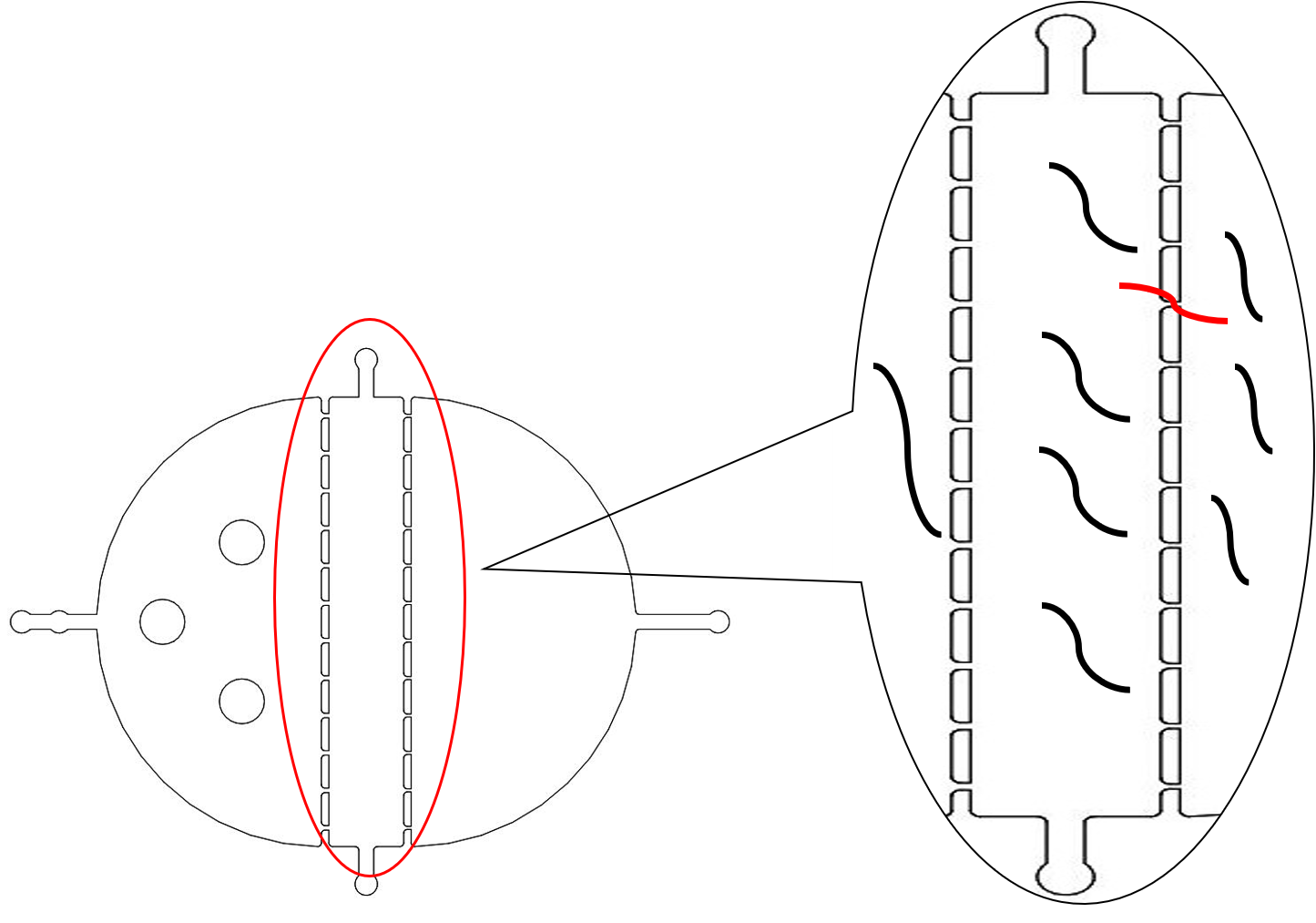
{{SUSTech_Image_Center_8 | filename=T--SUSTech_Shenzhen--Microfuildics--selective1.png|width=600px|caption=<B>Fig.2 The Problems of the Selective chip.</B> The efficiency of the chip was very low because of the small amount of the fences. Besides, the worms will also escape from the second fences just like the red one.}} | {{SUSTech_Image_Center_8 | filename=T--SUSTech_Shenzhen--Microfuildics--selective1.png|width=600px|caption=<B>Fig.2 The Problems of the Selective chip.</B> The efficiency of the chip was very low because of the small amount of the fences. Besides, the worms will also escape from the second fences just like the red one.}} | ||
| − | + | The second plan was the <i>C. elegans’</i> synchronization.(how to do the <i>C. elegans’</i> synchronization 链接到 protocl) We got the embryos (Fig.3) from the old worms so that the worms would be at the same stage because of the hatches of the embryos were at the same time. We selected several conditions of the synchronization, finally, we could get the worms at the same stage. The synchronous rate ({{SUSTech_Shenzhen/math|equ=<nowiki>\frac{the\, number\, of\, the\, worms\, at\, L4}{the\, number\, of\, all\, worms}</nowiki>}}*100%) could reach to about 80%. | |
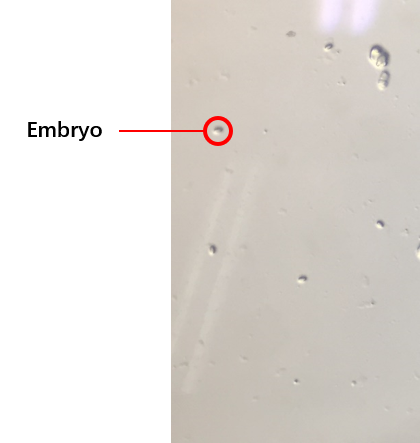
{{SUSTech_Image_Center_8 | filename=T--SUSTech_Shenzhen--2.png|width=600px|caption=<B>Fig.3 The Embryos of The Worms</B>}} | {{SUSTech_Image_Center_8 | filename=T--SUSTech_Shenzhen--2.png|width=600px|caption=<B>Fig.3 The Embryos of The Worms</B>}} | ||
| Line 41: | Line 41: | ||
== Gaussian Chip == | == Gaussian Chip == | ||
| − | The Gaussian chip (Fig.4) was designed to test if our exogenous genes | + | The Gaussian chip (Fig.4) was designed to test if our exogenous genes would influence their olfactory receptor neuron pair (preference and repulsion to some chemical odors). |
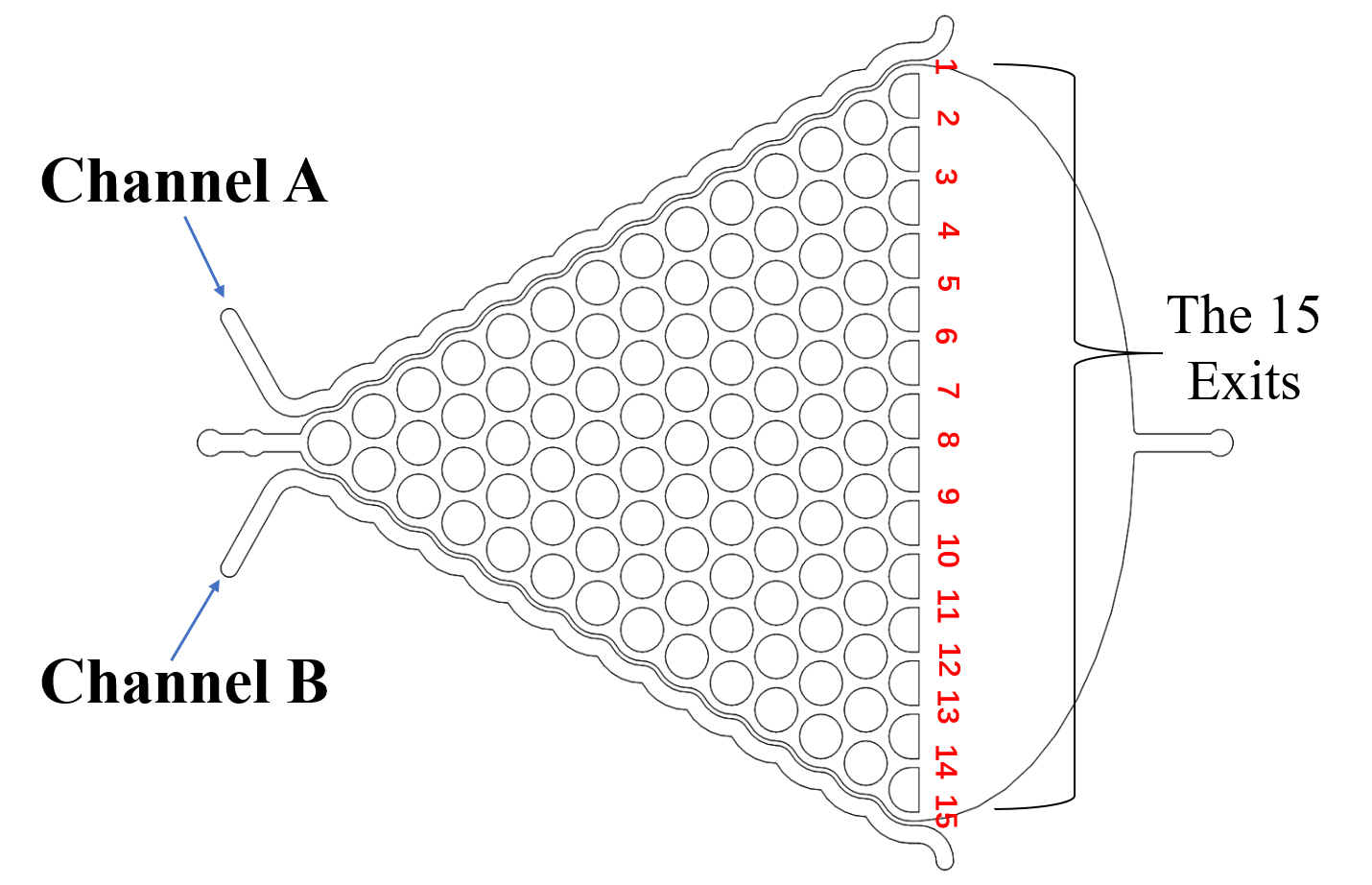
{{SUSTech_Image_Center_8 | filename=T--SUSTech_Shenzhen--Microfuildics--gstd.png|width=600px|caption=<B>Fig.4 The Gaussian Chip</B>}} | {{SUSTech_Image_Center_8 | filename=T--SUSTech_Shenzhen--Microfuildics--gstd.png|width=600px|caption=<B>Fig.4 The Gaussian Chip</B>}} | ||
| Line 51: | Line 51: | ||
{{SUSTech_Image_Center_8 | filename=T--SUSTech_Shenzhen--3.gif | caption=<B>Fig.6 The Experiment Process of the Gaussian Chip</B>}} | {{SUSTech_Image_Center_8 | filename=T--SUSTech_Shenzhen--3.gif | caption=<B>Fig.6 The Experiment Process of the Gaussian Chip</B>}} | ||
| + | |||
| + | The final result were not such a good Gaussian distribution like the Galton board because the <i>C. elegans'</i> choices were not absolutely normal. In order to adjust our results we built a model. https://2017.igem.org/Team:SUSTech_Shenzhen/Model3 | ||
== Immobilization Chip == | == Immobilization Chip == | ||
| Line 58: | Line 60: | ||
{{SUSTech_Image_Center_8 | filename=T--SUSTech_Shenzhen--9.png|width=600px|caption=<B>Fig.7 The Immobilization Chip</B>}} | {{SUSTech_Image_Center_8 | filename=T--SUSTech_Shenzhen--9.png|width=600px|caption=<B>Fig.7 The Immobilization Chip</B>}} | ||
| − | We could immobilize the worms in the worm traps (Fig.8) and watch the neuronal activity (Fig.9) successfully using fluorescence microscope (Nikon eclipse Ti) | + | We could immobilize the worms in the worm traps (Fig.8) and watch the neuronal activity (Fig.9) successfully using fluorescence microscope (Nikon eclipse Ti). |
{{SUSTech_Image_Center_8 | filename=T--SUSTech_Shenzhen--Microfuildics--guding1.jpeg|width=600px|caption=<B>Fig.8 The Worms in Immobilization Chip.</B>}} | {{SUSTech_Image_Center_8 | filename=T--SUSTech_Shenzhen--Microfuildics--guding1.jpeg|width=600px|caption=<B>Fig.8 The Worms in Immobilization Chip.</B>}} | ||
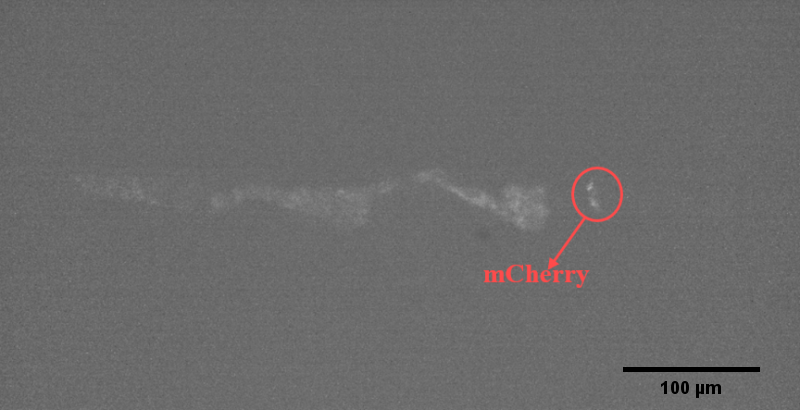
{{SUSTech_Image_Center_8 | filename=T--SUSTech_Shenzhen--Microfuildics--mcherry.png|width=600px|caption=<B>Fig.9 The mCherry of the AWA Neuro in the Odr10::CoChR::GEM-GECO::mCherry Worm.</B>}} | {{SUSTech_Image_Center_8 | filename=T--SUSTech_Shenzhen--Microfuildics--mcherry.png|width=600px|caption=<B>Fig.9 The mCherry of the AWA Neuro in the Odr10::CoChR::GEM-GECO::mCherry Worm.</B>}} | ||
| + | |||
| + | In addition, we could stimulate the Odr10::CoChR::GEM-GECO::mCherry worms to be active from the low state (Fig.10). On the other hand, we could also get the excitation wavelength of CoChR. The result showed that the light from the projector without filter (OD8) and the lights with 395 and 440 wavelengths from the LED of fluorescence microscope (Nikon eclipse Ti) could influence the C. elegans the other lights cannot. The lights with 395 and 440 wavelengths are closed to the ultraviolet which would hurt the C. elegans, so we got that the lights from the projector without the filter could active the CoChR. (参数啥的连接到光学) Unfortunately, we cannot see the clear neuro activity in str1::Chrimson::GEM-GECO::GFP worms using fluorescence microscope but we can use confocal microscope to observe the neuronal successfully. | ||
| + | |||
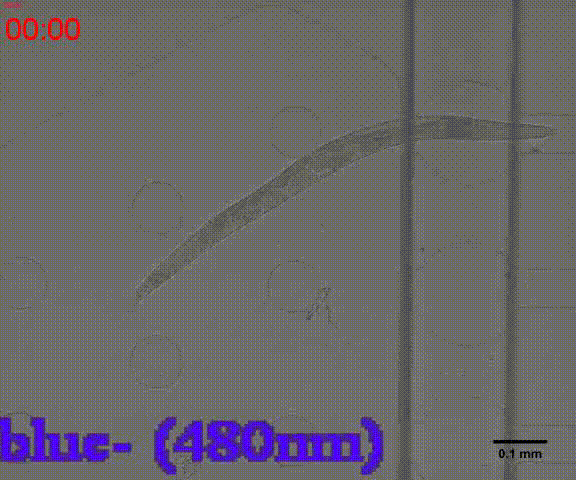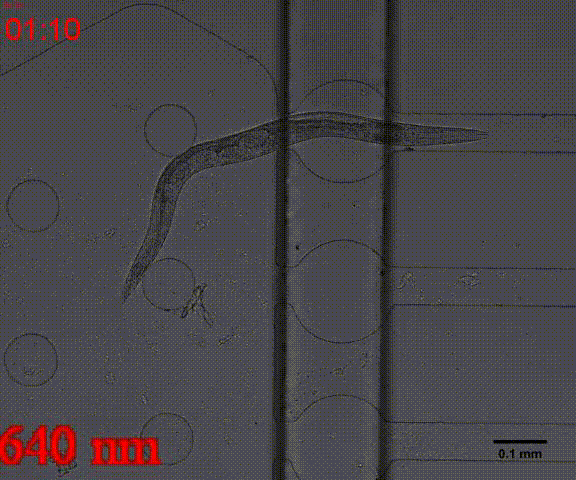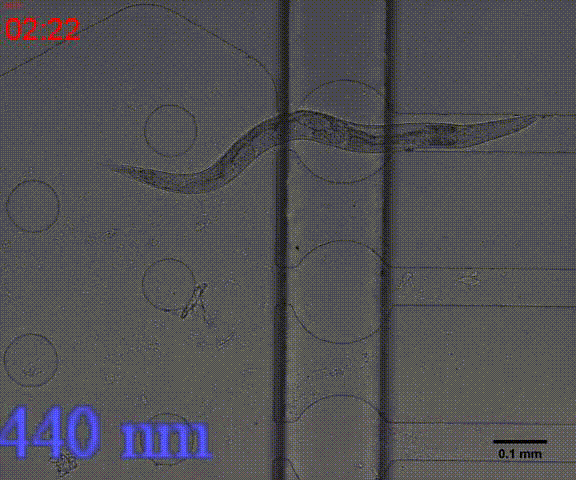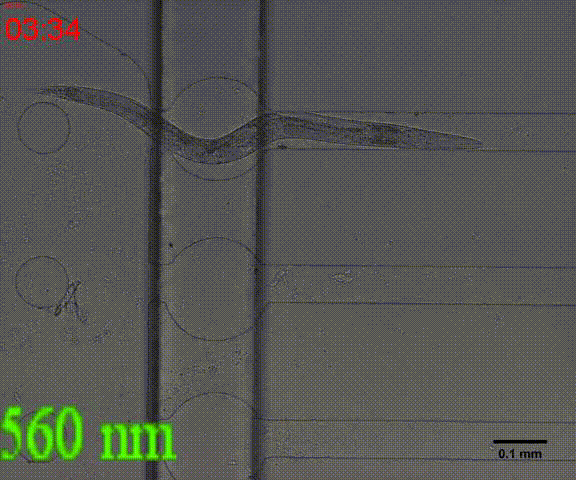
| + | {{SUSTech_Image_Center_8 | filename=T--SUSTech_Shenzhen--Microfuildics--wake up.gif|width=600px|caption=<B>Fig.10 The test of the CoChR using light with different wavelengths and intensity. </B>The lights <i>blue-</i> and <i>blue+</i> is from the projector whose wavelength is about 480 and the lights whose wavelengths are 395, 440, 470, 560 and 640 are from the LED of fluorescence microscope (Nikon eclipse Ti).}} | ||
= References = | = References = | ||
Revision as of 22:21, 25 October 2017
Results
Project
Contents
Optical Experiments Resutls
Under construction
| | First | Second | |--|------|---------| | 3 | 43 | 4 |
Microfluidic Experiments Resutls
Microfluidics were divide into three parts: the selective chip; the Gaussian chip; and the immobilization chip.
The Selection of the Caenorhabditis elegans
There are two plans of selecting worms. Why we need the C. elegans with the same stage The first one is using microfluidics. We designed the selective chip to select the Caenorhabditis elegans (C. elegans) with the appropriate size (Fig.1).
We need a large number of the worms with the same stage to do the Gaussion chip. Why we need so many with the same stage However, we found that the chip only has 12 fences (Fig.2) and the efficiency of the selective chip was very low because of such a small number of the fences. In addition, the C. elegans have flexible body, some of the suitable size worms would still go through the second fences (Fig.2).
The second plan was the C. elegans’ synchronization.(how to do the C. elegans’ synchronization 链接到 protocl) We got the embryos (Fig.3) from the old worms so that the worms would be at the same stage because of the hatches of the embryos were at the same time. We selected several conditions of the synchronization, finally, we could get the worms at the same stage. The synchronous rate (\frac{the\, number\, of\, the\, worms\, at\, L4}{the\, number\, of\, all\, worms}*100%) could reach to about 80%.
Gaussian Chip
The Gaussian chip (Fig.4) was designed to test if our exogenous genes would influence their olfactory receptor neuron pair (preference and repulsion to some chemical odors).
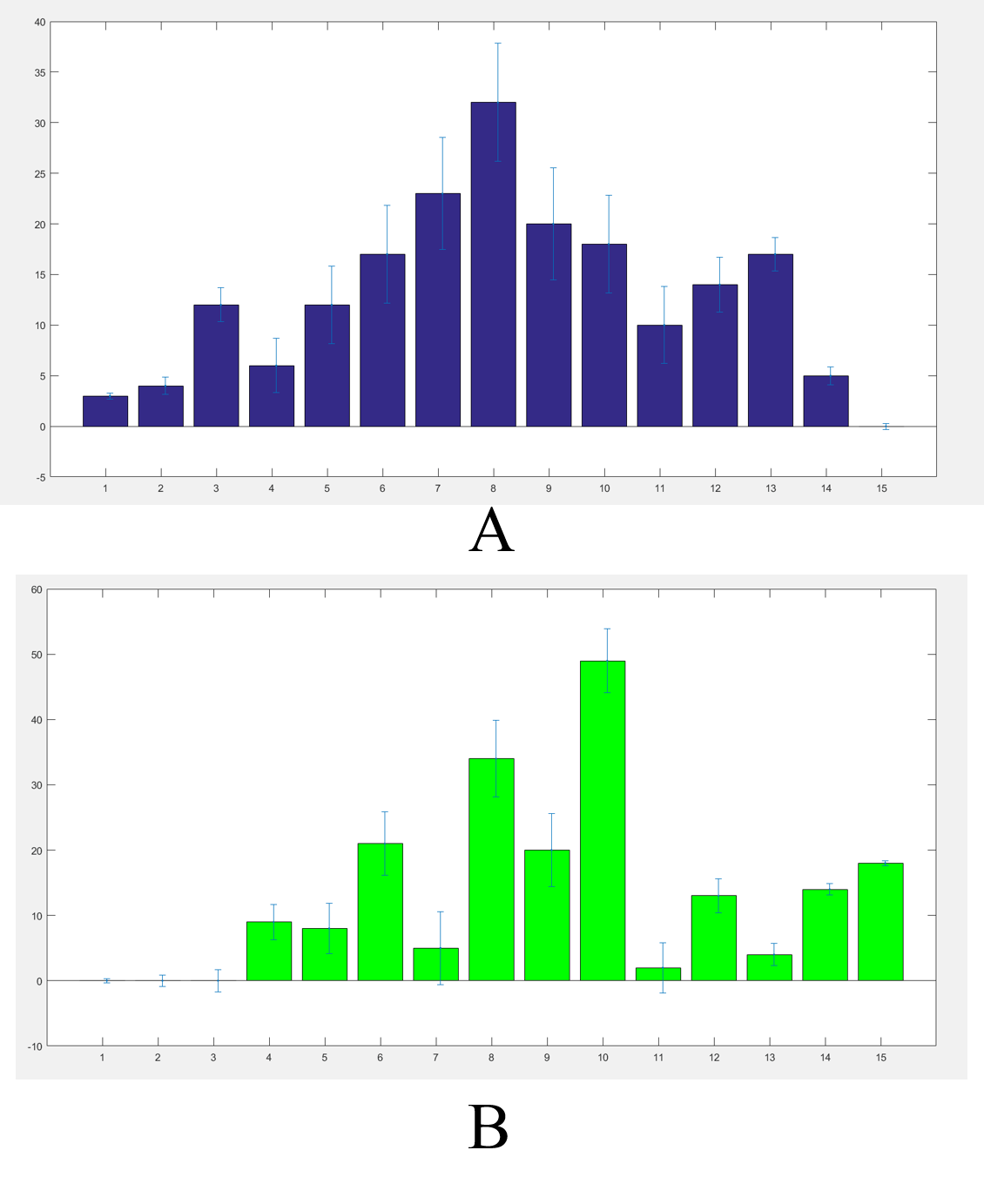
We got the worms’ distributions(Fig.5) after several experiments for the wildtype worms and our experimental worms with or without the chemicals (Fig.6).
The final result were not such a good Gaussian distribution like the Galton board because the C. elegans' choices were not absolutely normal. In order to adjust our results we built a model. https://2017.igem.org/Team:SUSTech_Shenzhen/Model3
Immobilization Chip
The immobilization chip was deigned to immobilize the C. elegans in worm traps or parallel channels for worm imaging and ethological experiments.
We could immobilize the worms in the worm traps (Fig.8) and watch the neuronal activity (Fig.9) successfully using fluorescence microscope (Nikon eclipse Ti).
In addition, we could stimulate the Odr10::CoChR::GEM-GECO::mCherry worms to be active from the low state (Fig.10). On the other hand, we could also get the excitation wavelength of CoChR. The result showed that the light from the projector without filter (OD8) and the lights with 395 and 440 wavelengths from the LED of fluorescence microscope (Nikon eclipse Ti) could influence the C. elegans the other lights cannot. The lights with 395 and 440 wavelengths are closed to the ultraviolet which would hurt the C. elegans, so we got that the lights from the projector without the filter could active the CoChR. (参数啥的连接到光学) Unfortunately, we cannot see the clear neuro activity in str1::Chrimson::GEM-GECO::GFP worms using fluorescence microscope but we can use confocal microscope to observe the neuronal successfully.
References