Brittneywick (Talk | contribs) |
Brittneywick (Talk | contribs) |
||
| Line 559: | Line 559: | ||
font-family: 'objektiv-mk1' !important; | font-family: 'objektiv-mk1' !important; | ||
font-size: 185% !important; | font-size: 185% !important; | ||
| − | font-weight: | + | font-weight: 400; |
} | } | ||
Revision as of 18:01, 29 October 2017

MODELING
Predict and optimize yield
Acetaminophen
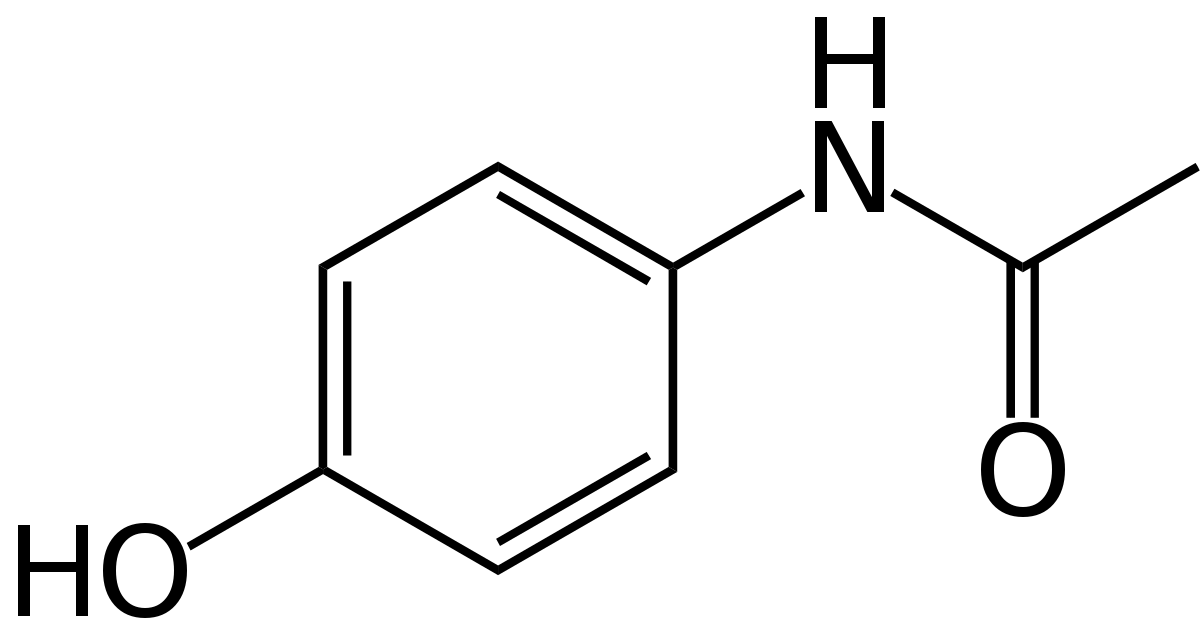
To predict acetaminophen biosynthesis, we analyzed the abundance of the acetaminophen's precursor, chorismate. Chorismate is a precursor primarily for the three aromatic amino acids, as well as salicylic acid, folate, vitamins, and many alkaloids [1]. We used published data on these products and gene transcription of amino acids for multiple species of cyanobacteria to assume the chorismate pool available to be processed into acetaminophen. Following that, we compared enzyme binding affinity Km's by creating a simulation of chorismate metabolism in Python to approximate enzyme rates. Insufficient data on cyanobacterial Km values necessitated using data from other bacterial species and comparing sequence identities to find the best match with our enzymes. We assumed the highest matching sequence Km's would be analagous to our enzyme's rate. With this information, we made a quantitative metabolic model which estimates how much of our precursor goes down each pathway.
We used different chorismate concentration estimates to reach several different estimates for acetaminophen, averaging 18.61mg ± 1.63mg acetaminophen per gram of Synechococcus biomass. This would be a sufficient quantity to detect through HPLC and serve as a starting point for optimizing production. This means that one 325mg dose of acetaminophen could be obtained in ~17g of biomass, meaning a 12 by 3 feet round pool could produce enough acetaminophen for more than 500 people every 10 days.
$$\frac{0.449\ mmol\ FWY}{1g\ biomass}\approx \frac{0.449\ mmol\ chor.}{1g\ biomass}\rightarrow\frac{1\ mol\ acet.}{3\ mol\ chor.}=\frac{0.15\ mmoles\ acet}{1\ g\ biomass}\times\frac{151.163g\ acet.}{1\ mol\ acet.}=\frac{22.62mg\ acet.}{1g\ biomass}$$To validate our organism's quantity of chorismate precursor, we used a custom Python program to convert DNA sequences to amino acids and calculate molar and mass percentages of chorismate derived aromatic amino acids. We ran both the genome and ribosomal protein sequences through our program, which resulted in 9.3% and 5.14% by mass aromatic amino acids. Using our higher sequence value of 9.3% and the assumption that our enzymes would take a third of the acetaminophen precursor, we estimate an acetaminophen concentration would be around 18mg per gram dried biomass.
$$\frac{0.093\ g\ FYW}{1\ g\ protein}\times\frac{0.6g protein}{1\ biomass}=\frac{0.056\ g\ FYW}{1g\ biomass}\rightarrow\frac{0.37\ mmol\ chor}{1\ g\ biomass}\times\frac{1\ mol\ acet}{3\ mol\ chor}\times\frac{151.163g\ acet}{1\ mol\ acet.}=\frac{18.61mg\ acet.}{1g\ biomass}$$$$\frac{0.051 g\ FYW}{1 g\ protein} * \frac{0.6 g\ protein}{1g\ biomass} = \frac{0.031g\ FWY}{1 g\ biomass} \rightarrow \frac{0.0.2973\ mmol\ chor}{1\ g\ biomass} * \frac{1\ mol\ acet}{3\ mol\ chor} *\frac{151.163\ g}{1 mol\ acet} = \frac{14.9\ mg\ acet}{1 g\ biomass}$$
These numbers show that there will probably be enough precursor to produce a useful, detectable quantity of acetaminophen. Based on literature and sequence estimates of aromatic amino acids, we can assume there would be at least that many moles of chorismate from which our added pathway pushes towards acetaminophen. The three calculations above can be averaged to finally predict 18.61mg ± 1.63mg acetaminophen per gram of Synechococcus biomass.
Vitamin B12

Predicting the quantity of active form of B12 produced depends on a successful metabolism of the active lower ligand, 5,6-dimethyl-benzimidizole (5,6-DMB) and the quantity of cobalt which binds to our engineered product. Cobalt is a limiting factor for growth in many marine environments, limiting formation of B12’s large corrin-ring. Even for species that use adenine to bind to the lower ligand and form pseudo-B12, cobalt supplementation enhances growth and thus is likely the limiting precursor for B12 production [5]. The gene BluB was inserted to enzymatically convert FMNH2 to our activating lower ligand 5,6-DMB. With this ligand available and our second gene insert, BluB, which preferentially attaches 5,6-DMB to the cobalt, the vast majority of our B12 should be made in active form. The gene that creates the lower ligand from the FMNH2, Ssue, came from Synechococcus elongatus 7002 and had the pTrc promoter rendering it a strong enough producer to prevent 5,6-DMB from being the limiting factor. Published HPLC results show that that Spirulina produces between 150-250µg pseudo-cobalamin per hundred grams dry weight with the non-human-usable adenine as the lower ligand [6]. If we assume our Blub/CobC enzyme complex works as well as it does in its origin organism, as assayed in Microbial Cell Factories paper [7], then research suggests 5,6-DMB has a much higher binding affinity for cobalt and thus nearly all of it will be converted to active-form B12 resulting in a production of almost exactly the Recommended Daily Value of 6µg B12 per 3g serving.
Biomass

To understand the production capacity of our organism, we aggregated growth data from published papers and all of our lab’s growth data. Using carrying-capacity-limited logistic growth curves to fit our data to an equation, we modelled dried biomass and cell count with respect to time. We have also used growth optimization papers [8,9] to add additional dependent variables of temperature, light intensity, and starter culture density to our equation.
Light Intensity: μE m-2 s-1
Temperature: ℃
Starting Density: g biomass/ L



