MattAFrench (Talk | contribs) |
|||
| (11 intermediate revisions by 3 users not shown) | |||
| Line 251: | Line 251: | ||
<div class="whiteboxmargin"> | <div class="whiteboxmargin"> | ||
<h1>Why</h1> | <h1>Why</h1> | ||
| − | <p> | + | <p>In our experiment, the production of differentially identifiable expression patterns is paramount. A library of promoters with robust expression patterns was required to achieve this. To further modulate the expression patterns, a promoter targeted short guide RNA (sgRNA)-Cas9 repression system was devised. These sgRNAs have a 20bp SEED region homologous to the promoter between the -10 and -35 box, a deactivated Cas9 associates with the sgRNA and is directed to the promoter through it’s homology, this will inhibit the RNA polymerase as it’s very large and bind with a high affinity. The strength of inhibition is proportional to the binding afinity of the SEED region. <br> |
| + | A two vector based system was decided upon to express this system. One low copy vector will express the catalytically inactive Cas9 protein, promoters and a downstream reporter; the other will contain the sgRNAs and a high copy origin of replication (ori). Confining the sgRNAs to a separate vector is advantageous in that it can be easily substituted out for another vector, potentially detailing sgRNAs with a differing affinity to the promoter. This interchangeability can facilitate a ‘plug and play’ system to produce a multitude of expression levels with ease. | ||
| + | </p> | ||
<h1>How</h1> | <h1>How</h1> | ||
| − | <p> Refer to step 2 for plasmid construction. | + | <p> Refer to step 2 for plasmid construction.To select for a dual transformation, each vector backbone requires separate antibiotic resistances. Consequently, the high copy vector (containing sgRNAs) was given an erythromycin resistant cassette, the low copy backbone was given a Chloramphenicol resistance cassette. Each vector also required a gram-negative compatible ori. High copy ColE1 was present on the sgRNA vector to drive high expression and promoter inhibition. A low copy pSC101 ori was initially given to the other, as dCas9 is toxic in high concentrations. |
| + | Another set of sgRNAs were designed to have no inhibitory activity, termed sgRNA0s. The ‘seeds’ were not homologous to any of the promoter, whereas the other set were 100% homologous. This will act as a control, to signify how the expression will differ when inhibition is occurring. It will also serve later to increase the pool of possible expression patterns when we come to the random ligation experiments. These were cloned using traditional type II restriction cloning with BsaI sites.<br><br> | ||
| + | To construct the sgRNAs, primer dimers were PCR amplified to generate the initial fragments of the sgRNA, it’s promoter, and terminator; also with their complimentary BsaI sites. After digestion with BsaI, the fragments were linearly ligated to produce promoter-sgRNA-terminator ‘bricks’. These were then amplified using a set of ‘universal’ primers designed to be capable of amplifying all of our ‘bricks’, these primers were complementary to a multiple cloning site (MCS) flanking each brick we produced. These sgRNA bricks were then digested with this MCS and ligated into their associated high copy backbone. | ||
| + | </p> | ||
<h1>Results</h1> | <h1>Results</h1> | ||
| − | <p> | + | <p>For the CRISPRi results check step 6 and the lab-book for more in detail descriptions of the prodution of the gRNA and gel images.</p> |
| Line 291: | Line 296: | ||
<p>After this, the same method as above was used for the final dCas9 including vector. However with an addition of dCas9 in the final ligation phase. In this way two promoter libraries were produced.</p> | <p>After this, the same method as above was used for the final dCas9 including vector. However with an addition of dCas9 in the final ligation phase. In this way two promoter libraries were produced.</p> | ||
<h1>Results</h1> | <h1>Results</h1> | ||
| − | <p> | + | <p>For results check steps 3 and 4.</p> |
| Line 316: | Line 321: | ||
</p> | </p> | ||
<h1>How</h1> | <h1>How</h1> | ||
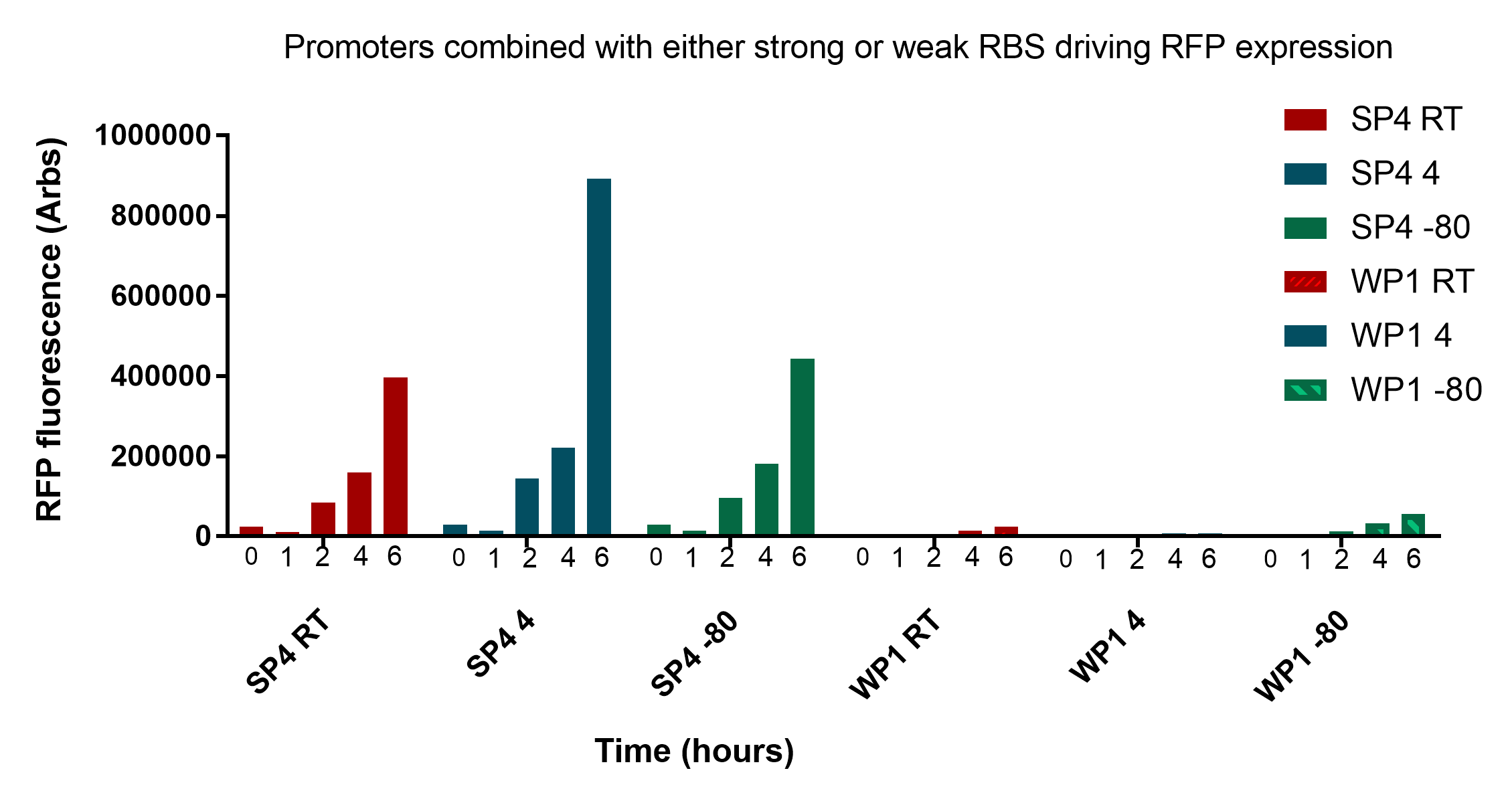
| − | <p> | + | <p>This growth curve was carried as described in week 15 of the lab book, please refer for full details. To summarise, overnight cultures of each specimen were inoculated into a new batch of media. These were then grown at 37°C for 24 hours, with measurements of OD600nm absorbance and fluorescence carried at 0, 2, 4, 6, and 24 hours. The promoters were paired with a downstream RFP reporter gene.</p> |
<h1>Results</h1> | <h1>Results</h1> | ||
| Line 414: | Line 419: | ||
<p>Interestingly, as predicted there is a substantial amount of fluorescence detectable 2 hours after revival, suggesting that a bacterial key could possibly be identifiable with little waiting period. However, they are undoubtedly more pronounced at 4 and 6 hours which would likely to be a more accurate time to measure.</p> | <p>Interestingly, as predicted there is a substantial amount of fluorescence detectable 2 hours after revival, suggesting that a bacterial key could possibly be identifiable with little waiting period. However, they are undoubtedly more pronounced at 4 and 6 hours which would likely to be a more accurate time to measure.</p> | ||
<p><table><tr><img src="https://static.igem.org/mediawiki/2017/f/f4/Groningen_data_revival.jpeg";></tr> | <p><table><tr><img src="https://static.igem.org/mediawiki/2017/f/f4/Groningen_data_revival.jpeg";></tr> | ||
| − | <tr><th> Figure 2. This graph represents data kindly given to us by the University of Groningen. It details the revival of the two samples containing strong RBS promoter 4 and weak RBS promoter 1 with a RFP reporter. These were grown for an 6 hour period and fluorescence measurements were taken at 0, 1, 2, 4, and 6 hours. One sample each was stored at either room temperature, 4<sup>o</sup>C, or -80<sup>o</sup>C.</th><tr></table></p> | + | <tr><th> Figure 2. This graph represents data kindly given to us by the University of Groningen. It details the revival of the two samples containing strong RBS promoter 4 and weak RBS promoter 1 with a RFP reporter. These were grown for an 6 hour period and fluorescence measurements were taken at 0, 1, 2, 4, and 6 hours. One sample each was stored at either room temperature, 4<sup>o</sup>C, or -80<sup>o</sup>C.</th><tr></table></p><br> |
<p> | <p> | ||
| − | In figure 2, samples were transported to the University of Groningen through the postal courier service, so the samples at -80<sup>o</sup>C would not have been at this temperature for the entire duration up until | + | In figure 2, samples were transported to the University of Groningen through the postal courier service, so the samples at -80<sup>o</sup>C would not have been at this temperature for the entire duration up until measuremment. With this in mind, it still appears what was discussed before still holds true. </p> |
<br> | <br> | ||
| − | <p>In conclusion, these results are very promising and show that cells remain viable and reliably express the reporter for up to three weeks at convenient storage conditions necessary for our key to work in a practical sense. </p> | + | <p>In conclusion, these results are very promising and show that cells remain viable and reliably express the reporter for up to three weeks at convenient storage conditions necessary for our key to work in a practical sense. The two promoters SP4 and WP1 are very distinct and easily discernible. There still remains a significant amount of viable bacteria regardless of storage conditions and their fluorescence intensity is very comparable at all stages of growth. It is worth noting that measurements were taken at 0 and 1 hours in figure 2, and these showed minimal fluorescent intensity, suggesting that further optimisation needs to occur to achieve 0-1 hour readings. For details on the revival protocol, download <a style="font-weight:bold;" href="https://static.igem.org/mediawiki/2017/9/9d/Revival_protocol.docx">here.</a></p> |
<br> | <br> | ||
<p>For future research, this process could be optimised to produce even more reproducible results and yield a more robust identification process. Distinct 0 hour expression levels might also be possible, greatly increasing the potential of <i>Key. coli</i> becoming a common security system. Multiple reporters should be the next focus to strengthen the security of the key. All reporters will need to be tested for their stability and reproducibility. </p> | <p>For future research, this process could be optimised to produce even more reproducible results and yield a more robust identification process. Distinct 0 hour expression levels might also be possible, greatly increasing the potential of <i>Key. coli</i> becoming a common security system. Multiple reporters should be the next focus to strengthen the security of the key. All reporters will need to be tested for their stability and reproducibility. </p> | ||
| + | <br> | ||
| + | <img src="https://static.igem.org/mediawiki/2017/9/91/T--UNOTT--GroningenData.png" style="width:100%;height:auto;"> | ||
| + | <p class="imgcaption"><tr><th><b>Figure 3:</b> Testing the affect of storage time at various storage temperatures (Room temperature, 4°C and -80°C) on the fluorescence of revived cells expressing strong or weak RFP expression</th></tr></table></p> | ||
<br> | <br> | ||
<br> | <br> | ||
Latest revision as of 03:59, 2 November 2017

EXPERIMENTS: