Successfully digested, ligated, and transformed rfbE into pSB1C3
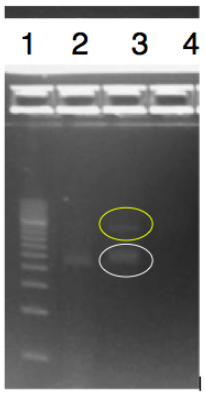
The objective of this experiment was to digest and ligate the rfbE BioBrick BBa_K2495001 into the pSB1C3 linearized backbone for DNA submission and characterization. The rfbE gene was obtained in the BioBrick format from IDT. A Modified restriction double digest and ligation protocol was used and can be found on our protocol page. We confirmed this by running PCR and a gel. Lanes were loaded as follows:
- 500 bp Bio-Rad ladder.
- Positive control of amplified DNA using PCR primers BBa_K2495019 and BBa_K2495020.
- Ligated product, with the amplified rfbE in white and the ligated product itself in yellow.
- Negative control of water with PCR primers.
Optimized the LAMP reaction to 20 minutes without lysis or centrifugation
The optimal conditions for cell lysis were previously investigated using DH5α E. coli transformed with pGLO plasmid (see biology lab notebook. The optimal cell lysis conditions for use with LAMP were tested using rfbE transformed into E. coli.
DH5α E. coli was transformed with rfbE using the pJET cloning vector. Additionally, naturally occurring E. coli was isolated from a sample of beef that was left at room temperature overnight. Miniprep (Qiagen) was used to isolated the plasmid DNA of these cells for the following experiments.
LAMP was run according to the instructions of the manufacturer (OptiGene) using primers targeted at rfbE designed with PrimerExplorer V5. The experimental conditions were:
- rfbE gene fragment (IDT) - positive control
- Plasmid DNA of rfbE in DH5α - negative control
- Plasmid DNA of rfbE in DH5α treated with 10 minutes of 95˚C heat lysis - negative control
- Plasmid DNA of rfbE in DH5α
- rfbE in DH5α suspended in TE buffer and treated with 10 minutes of 95˚C heat lysis
- rfbE in DH5α suspended in TE buffer (without heat treatment)
- Plasmid DNA of beef E. coli
- Plasmid DNA of beef E. coli treated with 10 minutes of 95˚C heat lysis
LAMP did not require heat lysis to show the typical smear of a positive result. The experiment also showed negative results for the naturally occurring E. coli. .The ladders were loaded as follows:
- 500 bp Bio-Rad ladder.
- Positive control of rfbE synthesized by IDT
- rfbE
- rfbE with heat lysis
- rfbEwithout heat lysis
- DH5α
- DH5α with heat
- DH5α without heat
- Beef E. coli with heat
- Beef E. coli without heat
Due to the results of this experiment, we envision that it will not be necessary to include a heat lysis step in our final version of the detection process for Shiga-toxigenic bacteria

Sensitive to 106 cells/mL, the FDA minimum infectious dose
To determine the limit of detection of our LAMP reactions, we ran it on serial dilutions of inoculated broth and monitored the progress using the Applied Biosystems SteponePlus Real-Time PCR System (Thermo Fisher, MA). Inoculated broth was diluted and run according to the optimized protocol described in our biology lab notebook. The dilutions ranged from 108 cells/mL to 101 cells/mL. Six serial dilutions of a previously miniprepped sample were used as positive controls. The negative control was TE buffer. The reaction was run on a total volume of 12.5µl. Duplicates were averaged and the fluorescence was plotted against time using Excel.
The results corresponded with a subsequent gel on the amplified products. The band intensity of the gels were in agreement with the relative fluorescence intensities visualized during the reaction. The ladders were loaded as follows:

- 500 bp Bio-Rad ladder.
- Positive control
- Lab broth, negative control
- ddH2O, negative control
- 108 cells/mL
- 107 cells/mL
- 105 cells/mL
- 104 cells/mL
- 103 cells/mL
- 102 cells/mL
- 101 cells/mL
While samples of lower concentrations were run, we could not draw any conclusive evidence from these as they did not differ significantly from the fluorescence intensity of the control. The fluorescence intensity of the broth with 108 cells/mL was significantly higher than the highest concentration of miniprepped sample (66 ng/mL). This does not correspond to the gel, in which lane 2 shows a stronger band than in lane 5. This may be due to the high concentration of cell debris that may have inhibited the reaction. Concentrations between 107 cells/mL and 106 cells/mL have decreasing fluorescence intensity and we are confident that our detection is accurate to this bacterial concentration.
We have found that using a larger reaction volume (25µl) does increase our sensitivity significantly, but due to shipping difficulties we were unable to obtain enough MasterMix to continue testing with a 25µl reaction volume. Instead, subsequent tests were tested with 12.5µl volume.
Based on these results, we determined that our limit of detection is 106 cells/mL.

The reaction products were also visualized under blue light and UV light at 302 nm. We can clearly visualize a gradient in the fluorescence from highest to lowest concentrations of DNA. All samples are clearly more fluorescent than the negative control.









 When developing the heating method, we found that it needed to satisfy the following properties:
When developing the heating method, we found that it needed to satisfy the following properties:




