| Line 3: | Line 3: | ||
| − | < | + | <html> |
| − | < | + | <script> |
| − | + | $(document).ready(function(){ | |
| + | // Add smooth scrolling to all links | ||
| + | $("a").on('click', function(event) { | ||
| − | + | // Make sure this.hash has a value before overriding default behavior | |
| − | + | if (this.hash !== "") { | |
| − | + | // Prevent default anchor click behavior | |
| − | + | event.preventDefault(); | |
| − | + | ||
| − | + | ||
| − | + | // Store hash | |
| + | var hash = this.hash; | ||
| − | + | // Using jQuery's animate() method to add smooth page scroll | |
| − | + | // The optional number (800) specifies the number of milliseconds it takes to scroll to the specified area | |
| − | + | $('html, body').animate({ | |
| − | + | scrollTop: $(hash).offset().top | |
| − | + | }, 800, function(){ | |
| + | |||
| + | // Add hash (#) to URL when done scrolling (default click behavior) | ||
| + | window.location.hash = hash; | ||
| + | }); | ||
| + | } // End if | ||
| + | }); | ||
| + | }); | ||
| + | </script> | ||
| + | |||
| + | div class = "column full_size_outer" id = "top"> | ||
| + | <div class = "column_full_size_inner"> | ||
| + | <p> | ||
| + | Click on one of the images below to learn more about our results! | ||
| + | </p> | ||
| + | <br> | ||
| + | <div class="column sixth_size"> | ||
| + | <a href="#section1"><img src="https://static.igem.org/mediawiki/2017/0/0b/T--Austin_UTexas--GABA.png" style="width:100%"> <p>GABA Production </p></a> | ||
</div> | </div> | ||
| − | <div class=" | + | <div class="column sixth_size"> |
| + | <a href="#section2"><img src="https://static.igem.org/mediawiki/2017/d/de/T--Austin_UTexas--PartPlasmidGGA.png" style="width:100%"><p>Part Plasmids </p></a> | ||
| + | </div> | ||
| − | <div class="column | + | <div class="column sixth_size"> |
| + | <a href="#section3"><img src="https://static.igem.org/mediawiki/2017/f/fd/T--Austin_UTexas--GGACassette.png" style="width:100%"><p>Cassette Assembly</p></a> | ||
| + | </div> | ||
| + | <div class="column sixth_size"> | ||
| + | <a href="#section4"><img src="https://static.igem.org/mediawiki/2017/9/95/T--Austin_UTexas--LPlantarumCirc.png" style="width:100%"><p><i>Lactobacillus plantarum</i></p></a> | ||
| + | </div> | ||
| − | < | + | <br> |
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| − | |||
| − | < | + | <div class="column full_size"> |
| − | < | + | <br> |
| − | < | + | <div style="background-color: lightcyan;padding:14px;"> |
| − | </ | + | <h2 style="font-family: verdana; font-size: 35px">Golden Gate Assembly</h2> |
| − | </ | + | <p style="font-family: verdana">Although bacteria can naturally synthesize GABA, we wanted to increase expression of the <i>gadB</i> gene and subsequently GABA production in order to imbue a more potent medicinal quality to our probiotic, with the idea that this GABA-overproducing probiotic can then be consumed by patients with bowel disorders or anxiety (1). Overexpression of the <i>gadB</i> gene will be accomplished by placing it under the control of either the P8 or P32 constitutive promoters from <i>Lactococcus lactis</i> (2). |
| + | To make our GABA-producing probiotic we first needed to assemble a GABA overexpression cassette plasmid using the Golden Gate assembly method. The intention here is that bacteria containing this GABA overexpression cassette plasmid should produce high levels of GABA. In short, Golden Gate Assembly is a new cloning method that allows for the creation of a multi-part DNA assembly (i.e. cassette plasmid) in a single reaction through the use of DNA parts containing specific, predefined suffixes and prefixes with recognition sites for Type IIs restriction enzymes (e.g. BsmBI and BsaI). The specificity of these suffixes and prefixes provides directionality of the desired DNA parts during the assembly process. For our purposes, we used the MoClo Yeast Tool Kit by John Dueber (3). | ||
| − | < | + | <br> |
| + | <br> | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | </ | + | <h2 style="font-family: verdana; font-size: 35px">Part Assembly</h2> |
| + | <br> | ||
| + | <p style="font-family: verdana">The first part of our Golden Gate assembly workflow was part assembly, in which the <i>gadB</i> gene and the P8/P32 promoters were individually cloned into the entry vector pYTK001 (Fig. 1). The <i>gadB</i> gene and P8/P32 promoter sequences contain flanking BsmBI sites that produce overhangs compatible with those cut by BsmBI in the entry vector pYTK001. Thus, BsmBI cloning should result in part plasmids containing the <i>gadB</i> gene and P8/P32 promoters set within the pYTK001 backbone. </p> | ||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | </html> | ||
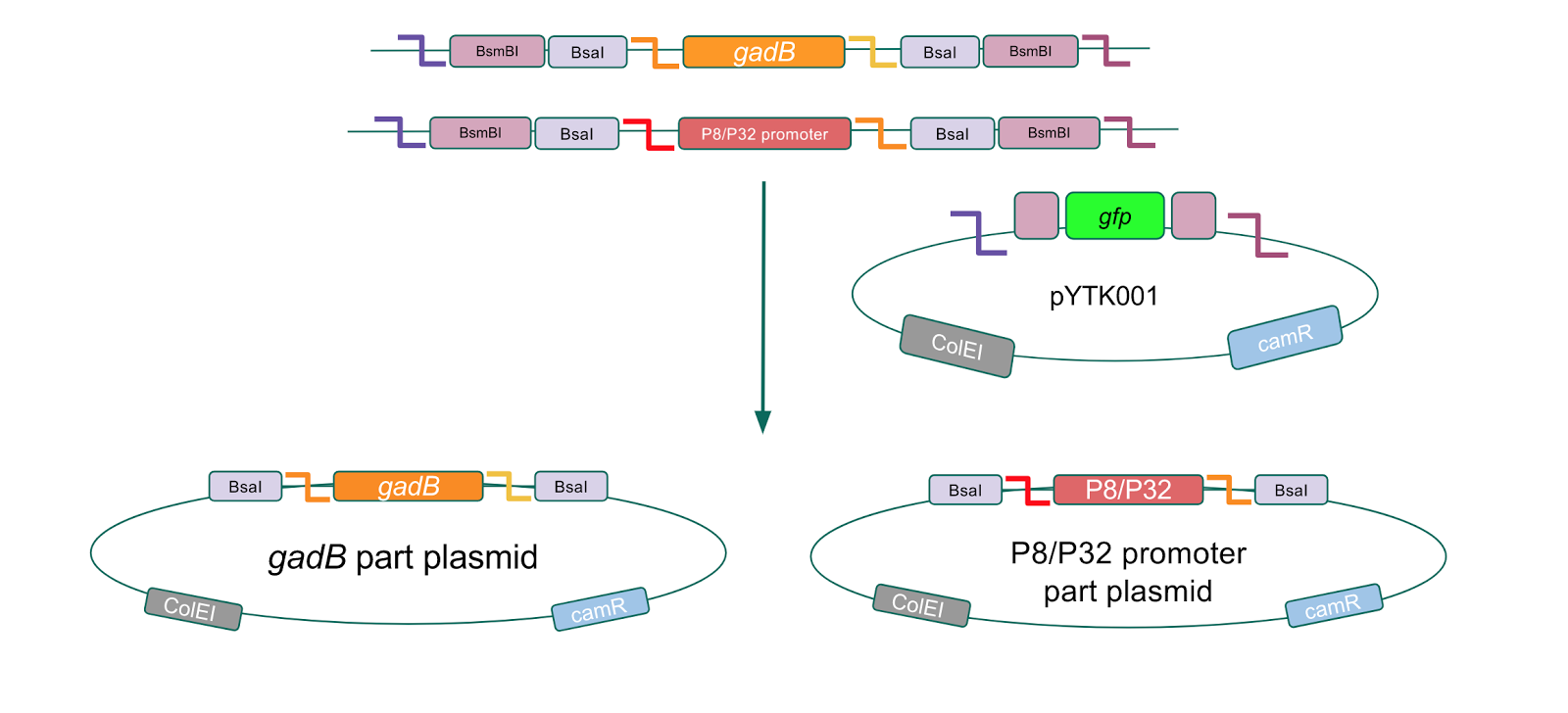
| + | [[File:T--Austin_UTexas--gadBAndPromoterPartPlasmids.jpg|thumb|center|800px|Figure 1. <i>gadB</i> gene and P8/P32 promoter part assembly process. Golden Gate compatible <i>gadB</i> and P8/P32 promoter sequences are cloned into the pYTK001 entry vector via BsmBI assembly.]] | ||
| + | <html> | ||
| + | |||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | <p style="font-family: verdana">Additionally, the BsmBI sites and overhangs in pYTK001 are flanking a <i>gfp</i> reporter gene. During the part assembly process, our DNA sequences of interest replaced this <i>gfp</i> reporter gene. This provided a phenotypic screen that allowed us to visually see which transformant colonies were negative and potentially positive. Under UV illumination, positive colonies containing our intended part plasmid assembly did not exhibit fluorescence under the UV illumination, while negative colonies did (Fig. 2.). The non-fluorescent colonies on the part plasmid transformation plates were miniprepped and subsequently sequence verified.</p> | ||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | </html> | ||
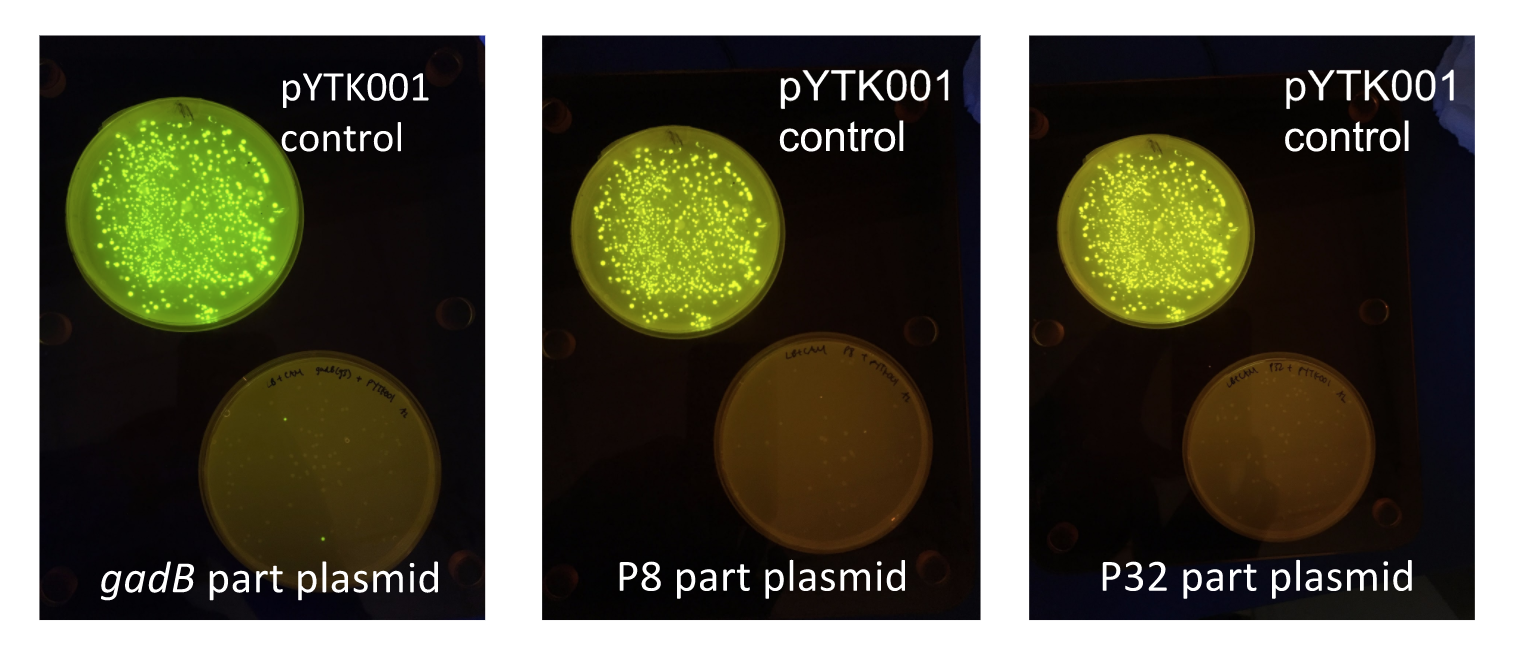
| + | [[File:T--Austin_UTexas--PartPlasmidTransformations.jpg|thumb|center|700px|Figure 2. <i>gadB gene</i> and P8/P32 promoter part plasmid <i>E. coli</i> transformations, compared to control transformations with pYTK001. Under UV illumination, transformants containing the correctly assembled part plasmids were non-fluorescent while negative transformants appeared fluorescent like colonies on the control plates.]] | ||
| + | <html> | ||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | |||
| + | <h2 style="font-family: verdana; font-size: 35px">Testing constitutive <i>Lactoccal</i> promoters </h2> | ||
| + | <p style="font-family: verdana">After successfully creating the <i>gadB</i> gene and P8/P32 promoter part plasmids, the functionality of these part plasmids were then assessed by assembling them into cassette plasmids.</p> | ||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | <p style="font-family: verdana">To test if our <i>Lactococcus lactis</i> constitutive promoters function well within <i>E. coli</i>, we created test cassette plasmids containing the <i>E2-Crimson</i> reporter gene (which encodes a red fluorescent protein) inserted downstream of either the P8 or P32 promoters using BsaI Golden Gate assembly. To create these test cassette, we used the P8/P32 promoter part plasmids, <i>E2-Crimson</i> part plasmid, an M13 terminator part plasmid, connector part plasmids, and pYTK095 vector as the backbone (Fig. 3). If the P8 and P32 promoters are functional in <i>E. coli</i>, we expected to observe red fluorescence In colonies transformed with our test cassette plasmids.</p> | ||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | </html> | ||
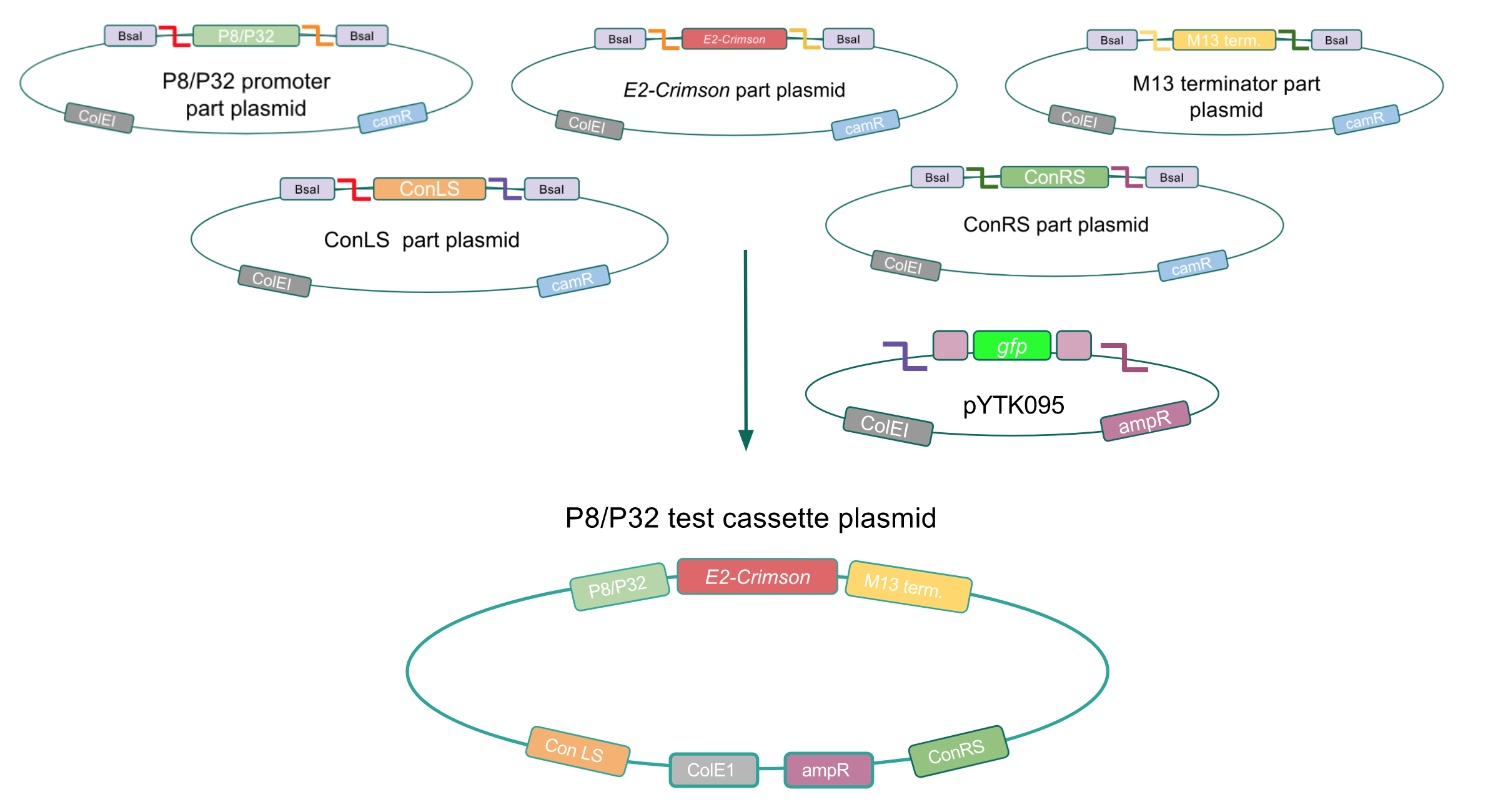
| + | [[File:T--Austin_UTexas--p8p32test.jpg|thumb|center|800px|Figure 3. Golden Gate assembly process of the P8 and P32 test cassette plasmids.]] | ||
| + | <html> | ||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | <p style="font-family: verdana">Upon first look, <i>E. coli</i> colonies transformed with these assemblies appeared purple-blue in color. This phenotype was due to the expression of the red fluorescent protein. Further, we noticed that <i>E. coli</i> colonies transformed with these assemblies fluoresced red under UV light, indicating that the P8 and P32 promoters are indeed expressing the <i>E2-Crimson</i> reporter gene and thus are functional in <i>E. coli</i> (Fig. 4).</p> | ||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | </html> | ||
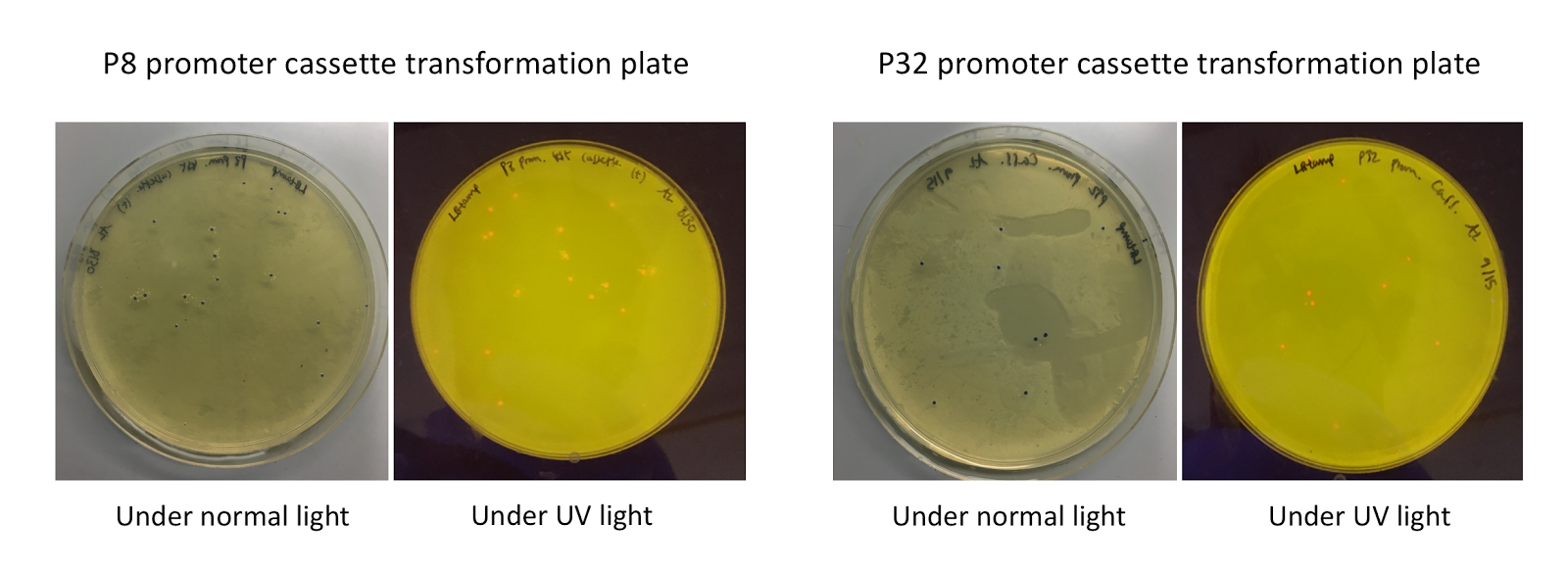
| + | [[File:T--Austin_UTexas--P8P32trans.jpg|thumb|center|800px|Figure 4. P8/P32 test cassette plasmid transformation plates, under normal and UV lights. Under normal lights, the colonies appeared purple-blue in color. Under UV, the colonies fluoresced red.]] | ||
| + | <html> | ||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | <h2 style="font-family: verdana; font-size: 35px">Testing for <i>gadB</i> overexpression in <i>E. coli</i></h2> | ||
| + | <p style="font-family: verdana">Using Golden Gate Assembly, we created cassette plasmids to test if the <i>gadB</i> gene could be overexpressed in <i>E. coli</i> via the P8 and P32 promoters. These cassette plasmids contained the <i>gadB</i> gene inserted downstream of either the P8 or P32 promoters. To make our <i>gadB</i> test cassette plasmids, we used pYTK095 as our backbone and part plasmids containing the P8/P32 promoters, <i>gadB</i> gene, M13 terminator, and connector sequences (Fig. 5).</p> | ||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | </html> | ||
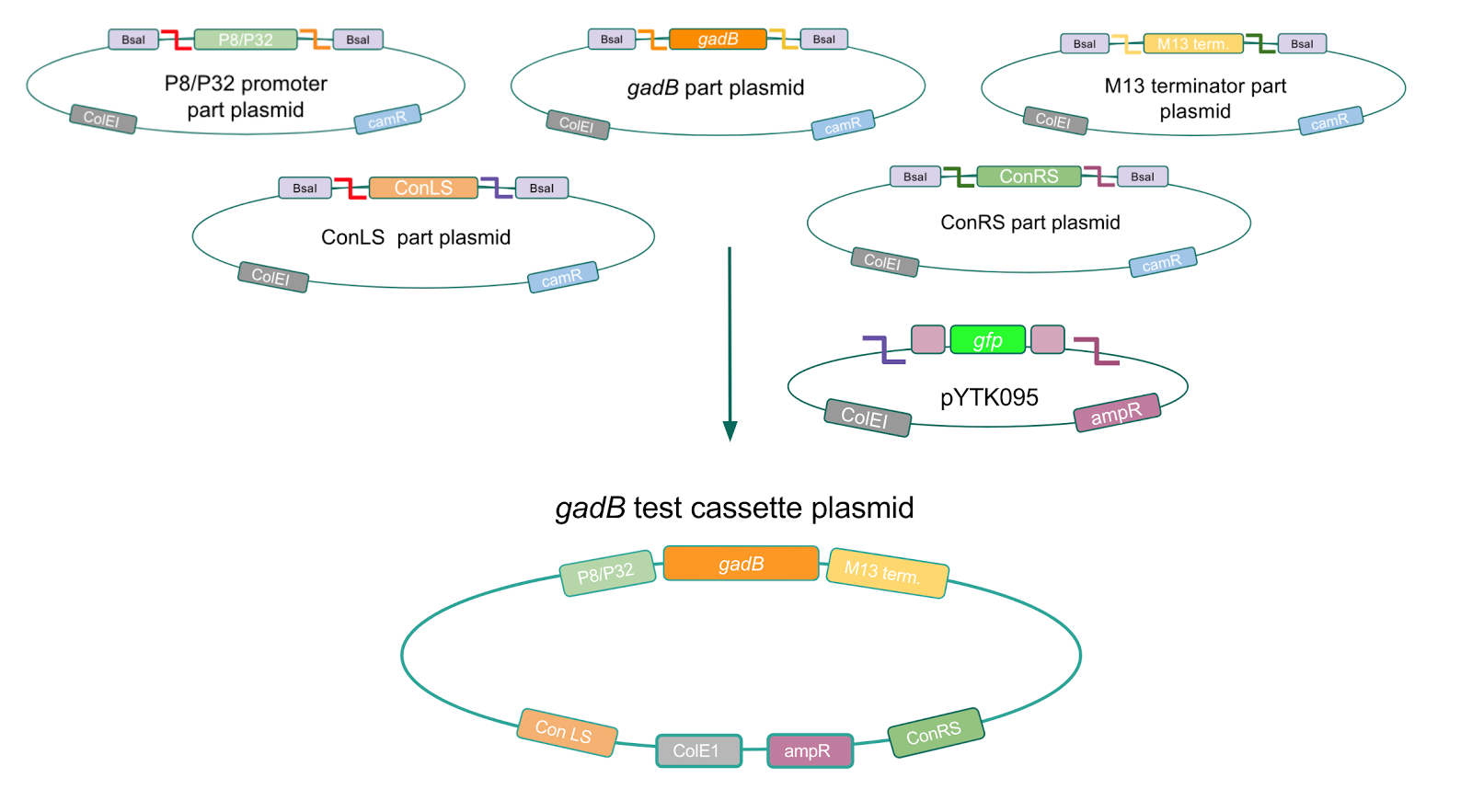
| + | [[File:T--Austin_UTexas--gadBTestCassette.jpg|thumb|center|800px|Figure 5. Golden Gate assembly process of P8/<i>gadB</i> and P32/<i>gadB</i> test cassette plasmids.]] | ||
| + | <html> | ||
| + | |||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | <p>Similar to the pYTK001 entry vector in part assembly, the pYTK095 backbone used for cassette assembly contained a <i>gfp</i> reporter gene that is replaced by sequences of interest. This allowed us to easily perform a phenotypic screen for positive colonies. Non-fluorescent colonies may potentially have had the correct cassette assembly, while fluorescent colonies did not (Fig. 6). The non-fluorescent colonies were then screened using colony PCR (Fig. 7). The positive colonies were then miniprepped and sequenced. The sequencing results indicated that in all of the samples there were several point mutations within the <i>gadB</i> gene. We hypothesized that overexpression of the <i>gadB</i> gene via the P8 and P32 constitutive promoters and the high-copy number ColE1 origin induced a high metabolic load on the cells, resulting in mutational degradation of <i>gadB</i>. To troubleshoot this problem, we decided to use a backbone containing the low-copy number p15A origin for the cassette assembly (Fig. 8)</p> | ||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | </html> | ||
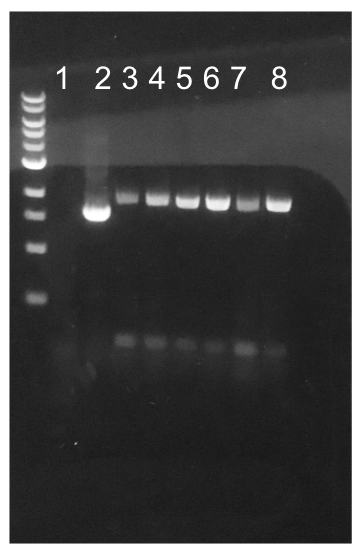
| + | [[File:T--Austin_UTexas--PromgadBgel.jpg|thumb|center|700px|Figure 7. Gel results for colony PCR of P8/<i>gadB</i> and P32/<i>gadB</i> non-fluorescent transformants. The primer pair used for this PCR experiment targeted the region of the cassette plasmid containing the DNA inserts (i.e. connectors, P8/P32 promoters, <i>gadB</i> gene, and M13 terminator). The expected band length was 2 kb. Lane 1 contains a negative PCR control. Lane 2 contains a positive PCR control. Lanes 3-5 contain the PCR products from three p8/<i>gadB</i> colonies. Lanes 6-8 contain the PCR products from three p8/<i>gadB</i> colonies. Evidently, all of the picked colonies were positive for the desired amplified region.]] | ||
| + | <html> | ||
| + | |||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | </html> | ||
| + | [[File:T--Austin_UTexas--TestCassetteLowCopy.jpg|thumb|center|800px|Figure 8. Modified Golden Gate assembly process of P8/<i>gadB</i> and P32/<i>gadB</i> test cassette plasmids using a backbone containing the low-copy number p15A origin.]] | ||
| + | <html> | ||
| + | |||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | |||
| + | <h2 style="font-family: verdana; font-size: 35px">Creating a Golden Gate compatible vector</h2> | ||
| + | |||
| + | <p style="font-family: verdana">After confirming <i>gadB</i> overexpression in <i>E. coli</i>, we want to assemble our final GABA overexpression cassette plasmid using the vector pMSP3535 as the backbone (Fig. 9). To do this, we first needed to make pMSP3535 Golden Gate compatible (i.e. free of BsaI restriction sites and containing correct overhangs for cassette assembly). We chose to work with pMSP3535 as it contains both a ColE1 origin for replication in <i>E. coli</i> and a pAMb1 origin for replication in Gram-positive bacteria including <i>Lactobacillus</i> species (4). Additionally, the pMSP3535 vector contains the resistance gene for erythromycin, which <i>Lactobacillus plantarum</i> is naturally susceptible to (5) | ||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | </html> | ||
| + | [[File:T--Austin_UTexas--Final_Cassette.jpg|thumb|center|800px|Figure 9. Golden Gate assembly of the GABA final overexpression cassette plasmid with the Golden Gate compatible pMSP3535 vector and the P8/P32 promoter, <i>gadB</i> gene, and M13 terminator part plasmids.]] | ||
| + | <html> | ||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
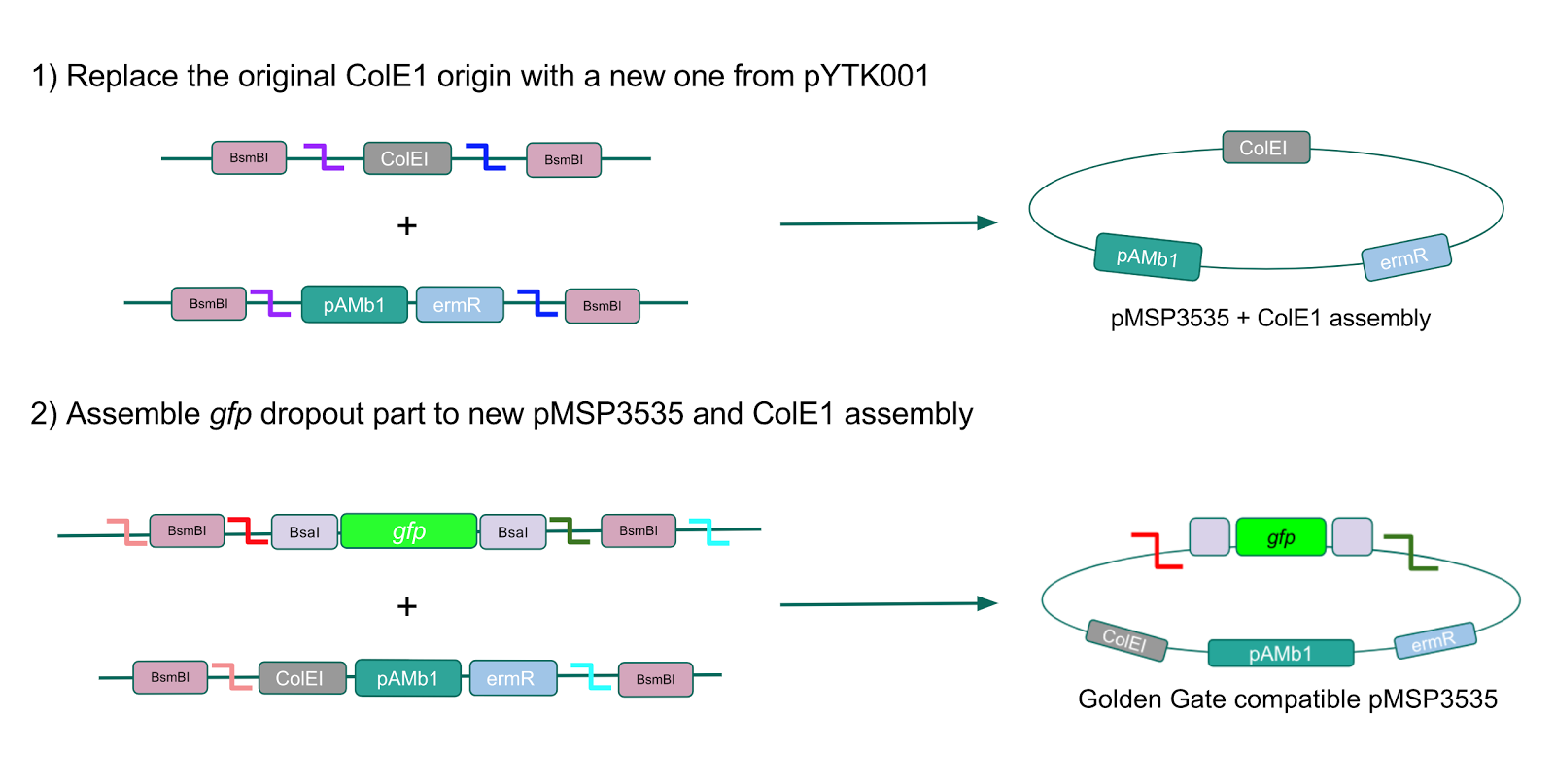
| + | <p style="font-family: verdana">The process of making the pMSP3535 vector Golden Gate compatible involved two steps: 1) assembling the pMSP3535 backbone (pAMb1 origin and erythromycin resistance gene) with a new ColE1 origin; 2) assembling a <i>gfp</i> dropout part to the assembly of the pMSP3535 backbone and the new ColE1 origin (Fig. 10).</p> | ||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | </html> | ||
| + | [[File:T--Austin_UTexas--CreatingpMSP3535GGA.jpg|thumb|center|800px|Figure 10. Assembly workflow for creating the Golden Gate compatible pMSP3535 backbone. The first step involves replacing the ColE1 origin in the pMSP3535 backbone with a new ColE1 origin from pYTK001 to create a pMSP3535 + ColE1 assembly. The second step involves combining the pMSP3535 + ColE1 assembly with a <i>gfp</i> dropout to form the final Golden Gate compatible vector to be used to create our GABA overexpression plasmid.]] | ||
| + | <html> | ||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | <p style="font-family: verdana">Since the original pMSP3535 vector contained two illegal BsaI sites within the ColE1 origin, we sought to replace this ColE1 origin with a BsaI-free one isolated from pYTK001. This assembly process involved linearizing and adding BsmBI sites and compatible overhangs to the pMSP3535 backbone and the pYTK001 ColE1 origin via PCR. After the pMSP3535 backbone and ColE1 origin were successfully amplified by PCR (Fig. 11a), they were joined together using BsmBI assembly. Diagnostic PCR was performed on pMSP3535 + ColE1 minipreps from <i>E. coli</i> transformants to screen for positive samples containing both the pMSP3535 backbone and the ColE1 inserts (Fig. 11b). After confirming the presence of the pMSP3535 vector and ColE1 origin, we partially sequence-confirmed the two miniprep samples.</p> | ||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | </html> | ||
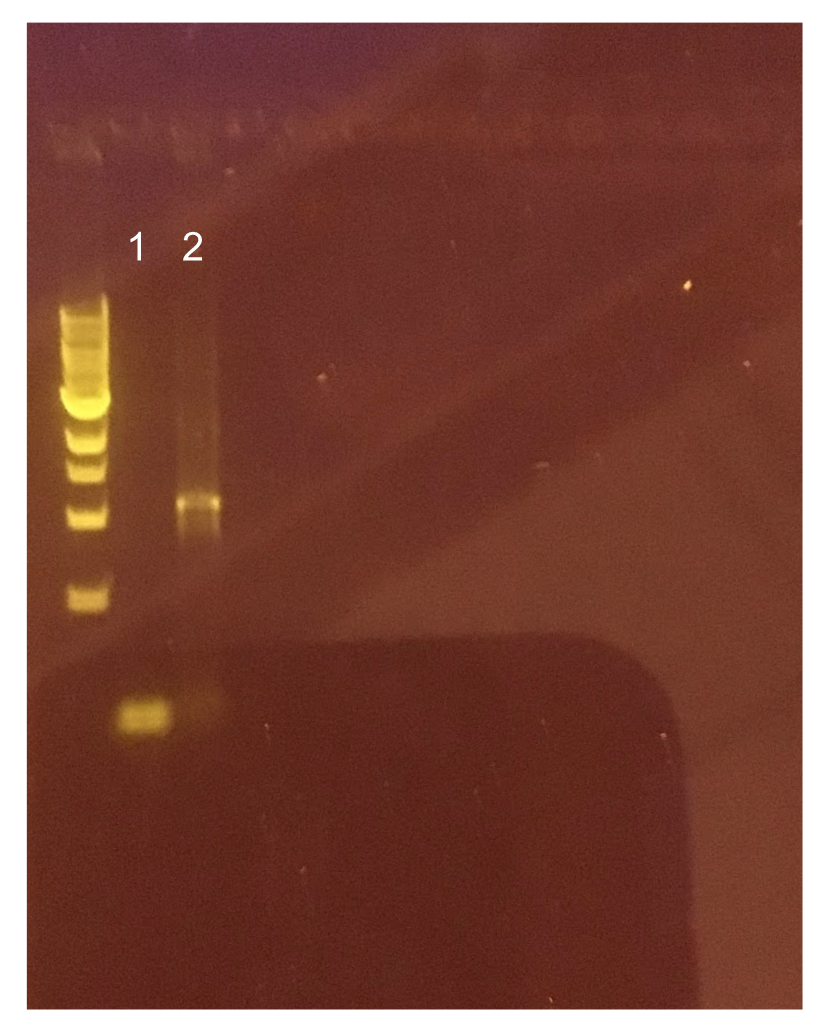
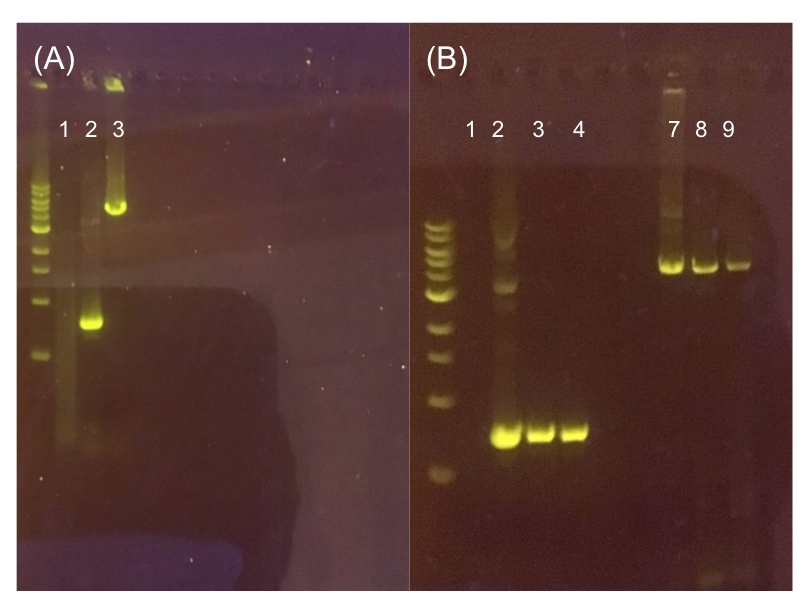
| + | [[File:T--Austin_UTexas--pMSP3535Gel.jpg|thumb|center|700px|Figure 11. (A) Agarose gel of successful PCR for the pMSP3535 backbone and the ColE1 origin. Lane 1 contains a negative PCR control. Lanes 2 and 3 contain the ColE1 origin (around 800 bp) and pMSP3535 backbone PCR products (around 4.3 kb). (B) Agarose gel of diagnostic PCR of miniprep samples from pMSP3535 + ColE1 transformants. Lane 1 contains a negative PCR control. Lane 2 contains a positive PCR control for the pMSP3535 backbone, while lanes 3 and 4 contain the pMSP3535 PCR from two miniprep samples. Lane 7 contains a positive PCR control for the ColE1 origin, while lanes 8 and 9 contain the ColE1 PCR from the same two miniprep samples. Evidently, the pMSP3535 backbone and ColE1 origin is present in both tested miniprep samples.]] | ||
| + | <html> | ||
| + | |||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | <p style="font-family: verdana">To this pMSP3535 + ColE1 assembly, we wanted to add a <i>gfp</i> dropout part containing internal BsaI sites that will generate overhangs compatible with those in the P8/P32 promoter and M13 terminator part plasmids. Additionally, the incorporation of this <i>gfp</i> dropout part will also allow us to visually screen for positive and negative transformants based on their fluorescence. BsmBI sites and compatible overhangs were added to the <i>gfp</i> dropout part by PCR amplifying it from pYTK047 (Fig. 12). We have been attempting to linearize and add BsmBI sites and overhangs to the positive pMSP3535 + ColE1 assemblies via PCR, with no success. However, results from diagnostic digests suggested that our assemblies may have contained extra, undesired DNA such as IS elements (Fig. 13). Thus, as of right now, we are screening for more positive pMSP3535 + ColE1 transformants. Once we have trouble-shooted this problem, the pMSP3535 + ColE1 and the <i>gfp</i> dropout PCR products will be joined through BsmBI assembly to form the final Golden Gate compatible pMSP3535 vector. | ||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | </html> | ||
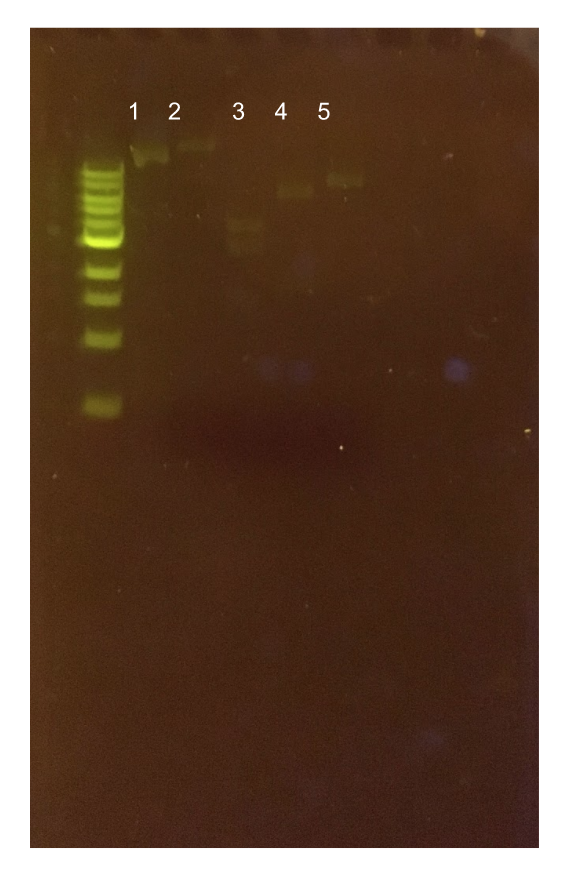
| + | [[File:T--Austin_UTexas--pYTK047Gel.jpg|thumb|center|700px|Figure 12. Agarose gel of <i>gfp</i> dropout PCR from pYTK047. Lane 1 contains a negative PCR control. Lane 2 contains the <i>gfp</i> dropout PCR product with an expected band length of around 1 kb.]] | ||
| + | <html> | ||
| + | |||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | </html> | ||
| + | [[File:T--Austin_UTexas--pMSP3535ColE1Diag.jpg|thumb|center|700px|Figure 13. Agarose gel of pMSP3535 + ColE1 assembly diagnostic digests. Lane 1 contains the undigested plasmid assembly. Lane 2 contains the ClaI-digested plasmid with expected 400 bp and 4.6 kb bands. The actual band generated is apparently above 10 kb. Lane 3 contains the XmnI-digested plasmid with expected 200 bp, 1.5 kb, and 3.3 kb bands. The generated band sizes were 2.5 kb and 4 kb. Lane 4 contains the KpnI-digested assembly with an expected 5.1 kb band. The generated band size was 6 kb. Lane 5 contains the Bg1II-digested assembly with an expected 5.1 kb band. The generated band size was 8 kb.]] | ||
| + | <html> | ||
| + | |||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | |||
| + | <h2 style="font-family: verdana; font-size: 35px">Assessing erythromycin susceptibility of <i>E. coli</i></h2> | ||
| + | <p style="font-family: verdana">Since we are creating our Golden Gate compatible pMSP3535 shuttle vector in <i>E. coli</i>, we wanted to determine the natural susceptibility of <i>E. coli</i> to erythromycin as the minimum concentration to use has not been established clearly in the literature. Thus, we performed an erythromycin minimum inhibitory concentration test in liquid LB media (Fig. 14). After one-day incubation, we observed that <i>E. coli</i> was resistant up to around 150 µg/mL of erythromycin. From this experiment, we have determined that the optimal erythromycin concentration for selecting against <i>E. coli</i> in liquid culture is around 200-250 µg/mL.</p> | ||
| + | |||
| + | <br> | ||
| + | <br> | ||
| + | |||
| + | </html> | ||
| + | [[File:T--Austin_UTexas--erymic.jpg|thumb|center|800px|Figure 14. Erythromycin minimum inhibitory concentration tests for <i>E. coli</i> in liquid media. From left to right: 0 µg/mL, 50 µg/mL, 100 µg/mL, 150 µg/mL, 200 µg/mL, 250 µg/mL, 300 µg/mL, 350 µg/mL, and 400 µg/mL.]] | ||
| + | <html> | ||
| + | |||
| + | |||
| + | <br> | ||
| + | |||
| + | |||
| + | |||
| + | <h2 style="font-family: verdana; font-size: 35px">References</h2> | ||
| + | <ol style="font-size:13px; font-family: verdana"> | ||
| + | |||
| + | <li>Yunes, R. A et al. GABA production and structure of <i>gadB</i>/<i>gadC</i> genes in <i>Lactobacillus</i> and <i>Bifidobacterium</i> strains from human microbiota. <i>Anaerobe</i>. 42: 197-204 (2016).</li> | ||
| + | |||
| + | <li>Zhu, D. et al. Isolation of strong constitutive promoters from <i>Lactococcus lactis</i> subsp. Lactis N8. <i>FEMS Microbiol Lett</i>. 363(16): pii: fnv107 (2015). </li> | ||
| + | |||
| + | <li>Lee, M. E. et al. A Highly Characterized Yeast Toolkit for Modular, Multipart Assembly. <i>ACS Synth. Biol</i>. 4(9): 975-86 (2015). </li> | ||
| + | |||
| + | <li>Pérez-Arellano, I. et al. Construction of Compatible Wide-Host-Range Shuttle Vectors for Lactic Acid Bacteria and <i>Escherichia coli</i>. Plasmid. 46(2): 106-16 (2001).</li> | ||
| + | |||
| + | <li>Flórez A. B. et al. Susceptibility of <i>Lactobacillus plantarum</i> Strains to Six Antibiotics and Definition of New Susceptibility–Resistance Cutoff Values. <i>Microbial Drug Resistance</i>. 12(4): 252-56 (2007).</li> | ||
| + | |||
| + | </ol> | ||
| + | |||
| + | |||
| + | |||
| + | </div> | ||
</html> | </html> | ||
Revision as of 22:52, 31 October 2017
Click on one of the images below to learn more about our results!
Golden Gate Assembly
Although bacteria can naturally synthesize GABA, we wanted to increase expression of the gadB gene and subsequently GABA production in order to imbue a more potent medicinal quality to our probiotic, with the idea that this GABA-overproducing probiotic can then be consumed by patients with bowel disorders or anxiety (1). Overexpression of the gadB gene will be accomplished by placing it under the control of either the P8 or P32 constitutive promoters from Lactococcus lactis (2).
To make our GABA-producing probiotic we first needed to assemble a GABA overexpression cassette plasmid using the Golden Gate assembly method. The intention here is that bacteria containing this GABA overexpression cassette plasmid should produce high levels of GABA. In short, Golden Gate Assembly is a new cloning method that allows for the creation of a multi-part DNA assembly (i.e. cassette plasmid) in a single reaction through the use of DNA parts containing specific, predefined suffixes and prefixes with recognition sites for Type IIs restriction enzymes (e.g. BsmBI and BsaI). The specificity of these suffixes and prefixes provides directionality of the desired DNA parts during the assembly process. For our purposes, we used the MoClo Yeast Tool Kit by John Dueber (3).
Part Assembly
The first part of our Golden Gate assembly workflow was part assembly, in which the gadB gene and the P8/P32 promoters were individually cloned into the entry vector pYTK001 (Fig. 1). The gadB gene and P8/P32 promoter sequences contain flanking BsmBI sites that produce overhangs compatible with those cut by BsmBI in the entry vector pYTK001. Thus, BsmBI cloning should result in part plasmids containing the gadB gene and P8/P32 promoters set within the pYTK001 backbone.
Additionally, the BsmBI sites and overhangs in pYTK001 are flanking a gfp reporter gene. During the part assembly process, our DNA sequences of interest replaced this gfp reporter gene. This provided a phenotypic screen that allowed us to visually see which transformant colonies were negative and potentially positive. Under UV illumination, positive colonies containing our intended part plasmid assembly did not exhibit fluorescence under the UV illumination, while negative colonies did (Fig. 2.). The non-fluorescent colonies on the part plasmid transformation plates were miniprepped and subsequently sequence verified.

Testing constitutive Lactoccal promoters
After successfully creating the gadB gene and P8/P32 promoter part plasmids, the functionality of these part plasmids were then assessed by assembling them into cassette plasmids.
To test if our Lactococcus lactis constitutive promoters function well within E. coli, we created test cassette plasmids containing the E2-Crimson reporter gene (which encodes a red fluorescent protein) inserted downstream of either the P8 or P32 promoters using BsaI Golden Gate assembly. To create these test cassette, we used the P8/P32 promoter part plasmids, E2-Crimson part plasmid, an M13 terminator part plasmid, connector part plasmids, and pYTK095 vector as the backbone (Fig. 3). If the P8 and P32 promoters are functional in E. coli, we expected to observe red fluorescence In colonies transformed with our test cassette plasmids.
Upon first look, E. coli colonies transformed with these assemblies appeared purple-blue in color. This phenotype was due to the expression of the red fluorescent protein. Further, we noticed that E. coli colonies transformed with these assemblies fluoresced red under UV light, indicating that the P8 and P32 promoters are indeed expressing the E2-Crimson reporter gene and thus are functional in E. coli (Fig. 4).
Testing for gadB overexpression in E. coli
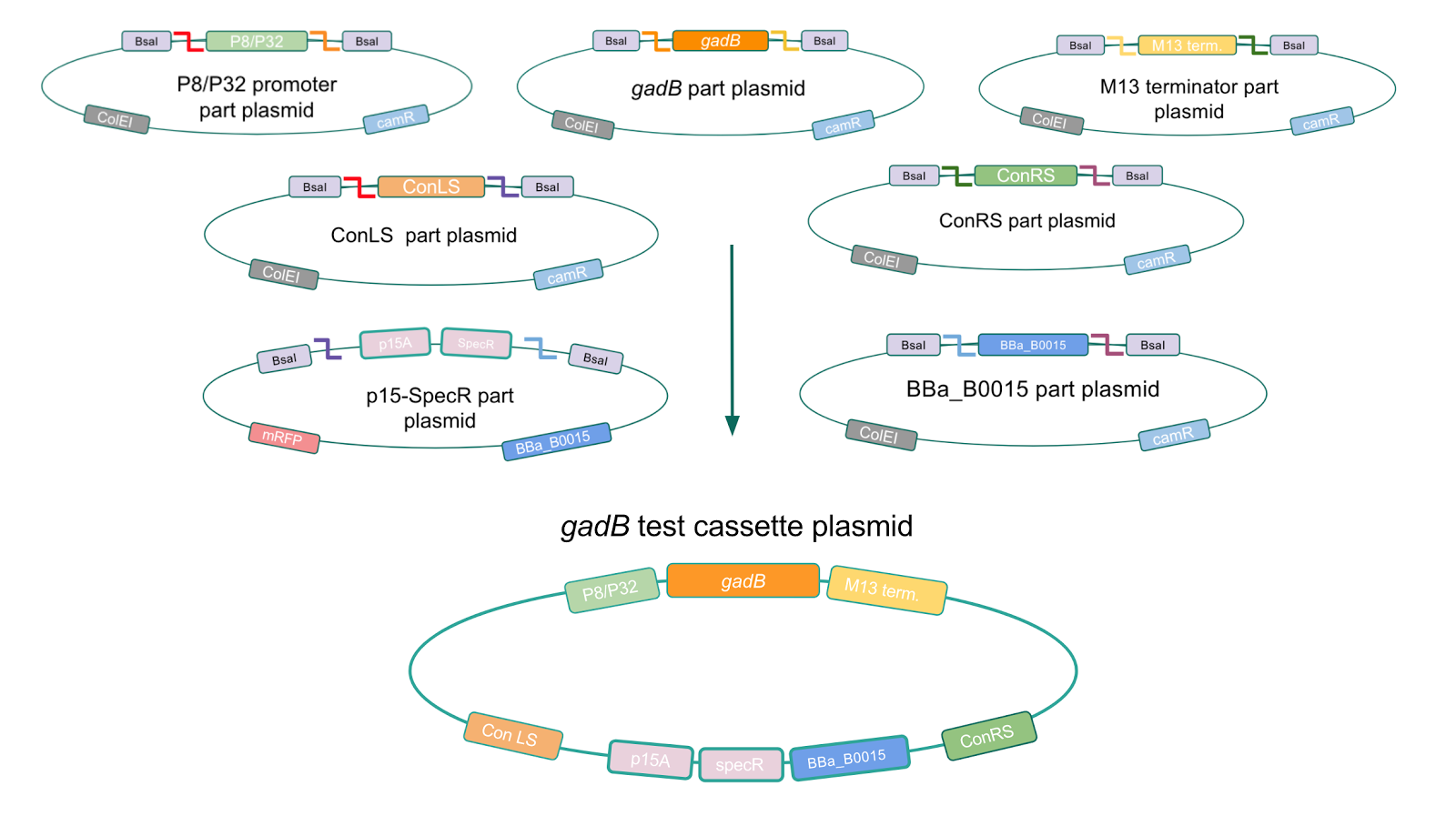
Using Golden Gate Assembly, we created cassette plasmids to test if the gadB gene could be overexpressed in E. coli via the P8 and P32 promoters. These cassette plasmids contained the gadB gene inserted downstream of either the P8 or P32 promoters. To make our gadB test cassette plasmids, we used pYTK095 as our backbone and part plasmids containing the P8/P32 promoters, gadB gene, M13 terminator, and connector sequences (Fig. 5).
Similar to the pYTK001 entry vector in part assembly, the pYTK095 backbone used for cassette assembly contained a gfp reporter gene that is replaced by sequences of interest. This allowed us to easily perform a phenotypic screen for positive colonies. Non-fluorescent colonies may potentially have had the correct cassette assembly, while fluorescent colonies did not (Fig. 6). The non-fluorescent colonies were then screened using colony PCR (Fig. 7). The positive colonies were then miniprepped and sequenced. The sequencing results indicated that in all of the samples there were several point mutations within the gadB gene. We hypothesized that overexpression of the gadB gene via the P8 and P32 constitutive promoters and the high-copy number ColE1 origin induced a high metabolic load on the cells, resulting in mutational degradation of gadB. To troubleshoot this problem, we decided to use a backbone containing the low-copy number p15A origin for the cassette assembly (Fig. 8)
Creating a Golden Gate compatible vector
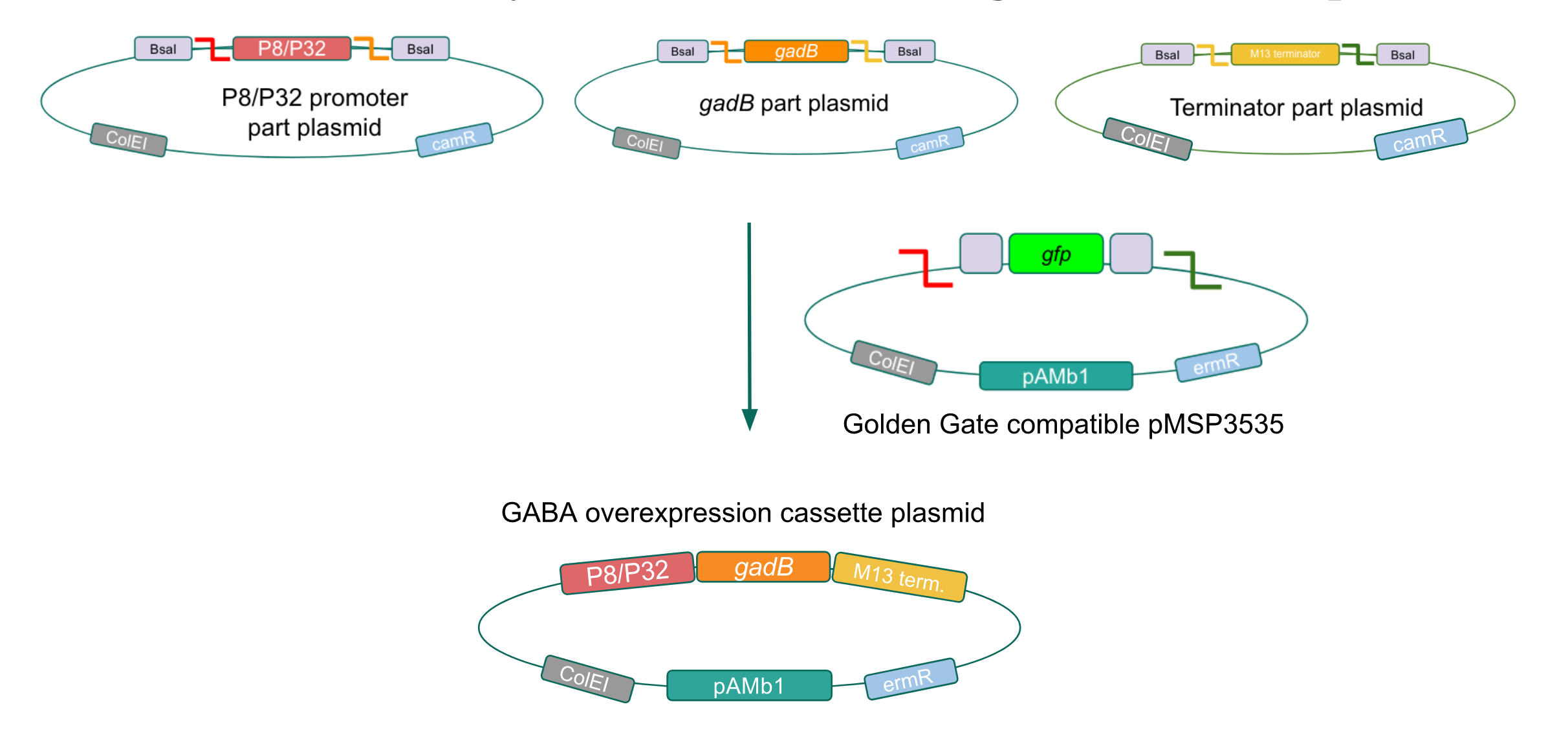
After confirming gadB overexpression in E. coli, we want to assemble our final GABA overexpression cassette plasmid using the vector pMSP3535 as the backbone (Fig. 9). To do this, we first needed to make pMSP3535 Golden Gate compatible (i.e. free of BsaI restriction sites and containing correct overhangs for cassette assembly). We chose to work with pMSP3535 as it contains both a ColE1 origin for replication in E. coli and a pAMb1 origin for replication in Gram-positive bacteria including Lactobacillus species (4). Additionally, the pMSP3535 vector contains the resistance gene for erythromycin, which Lactobacillus plantarum is naturally susceptible to (5)
The process of making the pMSP3535 vector Golden Gate compatible involved two steps: 1) assembling the pMSP3535 backbone (pAMb1 origin and erythromycin resistance gene) with a new ColE1 origin; 2) assembling a gfp dropout part to the assembly of the pMSP3535 backbone and the new ColE1 origin (Fig. 10).
Since the original pMSP3535 vector contained two illegal BsaI sites within the ColE1 origin, we sought to replace this ColE1 origin with a BsaI-free one isolated from pYTK001. This assembly process involved linearizing and adding BsmBI sites and compatible overhangs to the pMSP3535 backbone and the pYTK001 ColE1 origin via PCR. After the pMSP3535 backbone and ColE1 origin were successfully amplified by PCR (Fig. 11a), they were joined together using BsmBI assembly. Diagnostic PCR was performed on pMSP3535 + ColE1 minipreps from E. coli transformants to screen for positive samples containing both the pMSP3535 backbone and the ColE1 inserts (Fig. 11b). After confirming the presence of the pMSP3535 vector and ColE1 origin, we partially sequence-confirmed the two miniprep samples.
To this pMSP3535 + ColE1 assembly, we wanted to add a gfp dropout part containing internal BsaI sites that will generate overhangs compatible with those in the P8/P32 promoter and M13 terminator part plasmids. Additionally, the incorporation of this gfp dropout part will also allow us to visually screen for positive and negative transformants based on their fluorescence. BsmBI sites and compatible overhangs were added to the gfp dropout part by PCR amplifying it from pYTK047 (Fig. 12). We have been attempting to linearize and add BsmBI sites and overhangs to the positive pMSP3535 + ColE1 assemblies via PCR, with no success. However, results from diagnostic digests suggested that our assemblies may have contained extra, undesired DNA such as IS elements (Fig. 13). Thus, as of right now, we are screening for more positive pMSP3535 + ColE1 transformants. Once we have trouble-shooted this problem, the pMSP3535 + ColE1 and the gfp dropout PCR products will be joined through BsmBI assembly to form the final Golden Gate compatible pMSP3535 vector.
Assessing erythromycin susceptibility of E. coli
Since we are creating our Golden Gate compatible pMSP3535 shuttle vector in E. coli, we wanted to determine the natural susceptibility of E. coli to erythromycin as the minimum concentration to use has not been established clearly in the literature. Thus, we performed an erythromycin minimum inhibitory concentration test in liquid LB media (Fig. 14). After one-day incubation, we observed that E. coli was resistant up to around 150 µg/mL of erythromycin. From this experiment, we have determined that the optimal erythromycin concentration for selecting against E. coli in liquid culture is around 200-250 µg/mL.
References
- Yunes, R. A et al. GABA production and structure of gadB/gadC genes in Lactobacillus and Bifidobacterium strains from human microbiota. Anaerobe. 42: 197-204 (2016).
- Zhu, D. et al. Isolation of strong constitutive promoters from Lactococcus lactis subsp. Lactis N8. FEMS Microbiol Lett. 363(16): pii: fnv107 (2015).
- Lee, M. E. et al. A Highly Characterized Yeast Toolkit for Modular, Multipart Assembly. ACS Synth. Biol. 4(9): 975-86 (2015).
- Pérez-Arellano, I. et al. Construction of Compatible Wide-Host-Range Shuttle Vectors for Lactic Acid Bacteria and Escherichia coli. Plasmid. 46(2): 106-16 (2001).
- Flórez A. B. et al. Susceptibility of Lactobacillus plantarum Strains to Six Antibiotics and Definition of New Susceptibility–Resistance Cutoff Values. Microbial Drug Resistance. 12(4): 252-56 (2007).