| Line 26: | Line 26: | ||
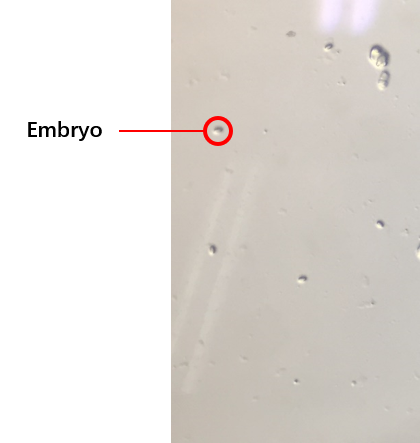
{{SUSTech_Image_Center_8 | filename=T--SUSTech_Shenzhen--2.png | caption=<B>Fig. 2 The Embryos after Bleaching Adults</B>}} | {{SUSTech_Image_Center_8 | filename=T--SUSTech_Shenzhen--2.png | caption=<B>Fig. 2 The Embryos after Bleaching Adults</B>}} | ||
| − | Compared with those two methods, we think the synchronization method is better. You can get more detail results by clicking <html><a target="_blank" href="https://2017.igem.org/Team:SUSTech_Shenzhen/Results/Microfluidic" class="btn btn-default"><i class="ion-arrow-right-c"></i> See Details</a></html>. | + | Compared with those two methods in experiment, we find that we can get almost 150 of 100 worms (One adult have no less than 3 embryos) in three days by the synchronization method, while we can just get 20 of 100 worms in one day by microfluidics method. Given that we need a large number of worms to do the following experiment, we think the synchronization method is better. You can get more detail results by clicking <html><a target="_blank" href="https://2017.igem.org/Team:SUSTech_Shenzhen/Results/Microfluidic" class="btn btn-default"><i class="ion-arrow-right-c"></i> See Details</a></html>. |
==2 The Gaussian Chip == | ==2 The Gaussian Chip == | ||
Revision as of 09:42, 30 October 2017
Microfluidics
Create for wisdom of Life
1 The Selection Chip
Most researches about Caenorhabditis elegans show that inserted genes can express well in worms at L4 stages. Thus, we need to select appropriate stages of worms to get the best experimental results. The simple method is to distinguish them by sizes, because worms in L4 stage have medium sizes. There are two plans of selecting worms. The first one is using microfluidics. With the flow contained worms going through this chip, only the medium sized worms can remain in the medium chamber, and we could collect them by injecting the flow from the bottom and gather them in the top. (Fig. 1)
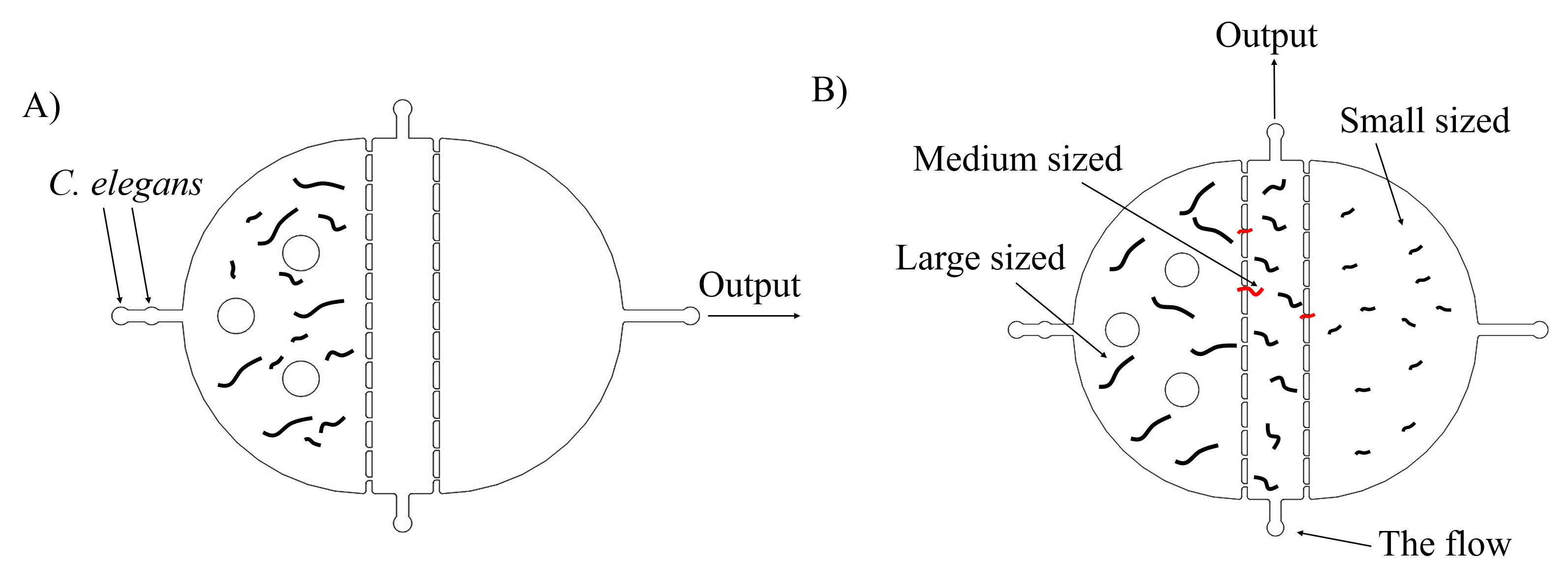
The second plan is making the grows of worms synchronously, which is utilized to get a large number of worms at the same stage (链接到微流控protocol). After collecting embryos (Fig. 2) by bleaching adults, we culture them and get a large number of worms at the same stages after three days. our synchronous rate is calculated as the formula below.
\frac{(the\,number\,of\, the\,worms\,at\,L4)*100}{the\, number\, of\, all\, worms}
The successful rate can reach to about 80%.
Compared with those two methods in experiment, we find that we can get almost 150 of 100 worms (One adult have no less than 3 embryos) in three days by the synchronization method, while we can just get 20 of 100 worms in one day by microfluidics method. Given that we need a large number of worms to do the following experiment, we think the synchronization method is better. You can get more detail results by clicking See Details.
2 The Gaussian Chip
In order to study locomotive behavior of C. elegans populations, we design the Gaussian Chip, a pillar-filled area, where the pillars are designed such that it allows crawling-like behaviors even though worms are immersed in liquid environment.(Fig. 3) [1].
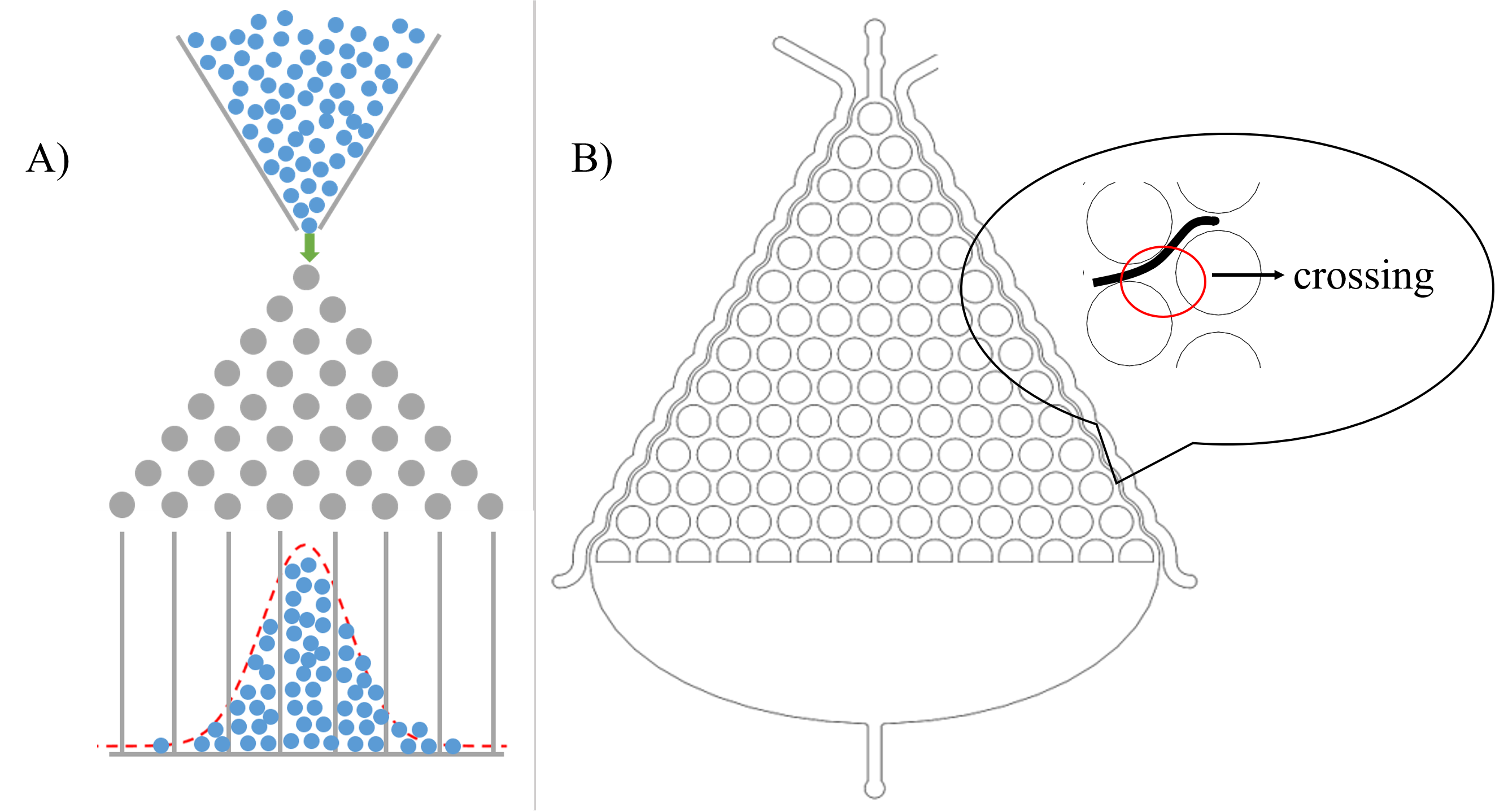
After deciding to use the microfluidics to study the locomotive behavior, we are noticed that the shape of microfluidics is similar to the Galton board.[2]. (Fig. 4(A)) Therefore, we assume that the probability for C. elegans choose to go left or right is equal when it passes a crossing. (Fig. 4(B))
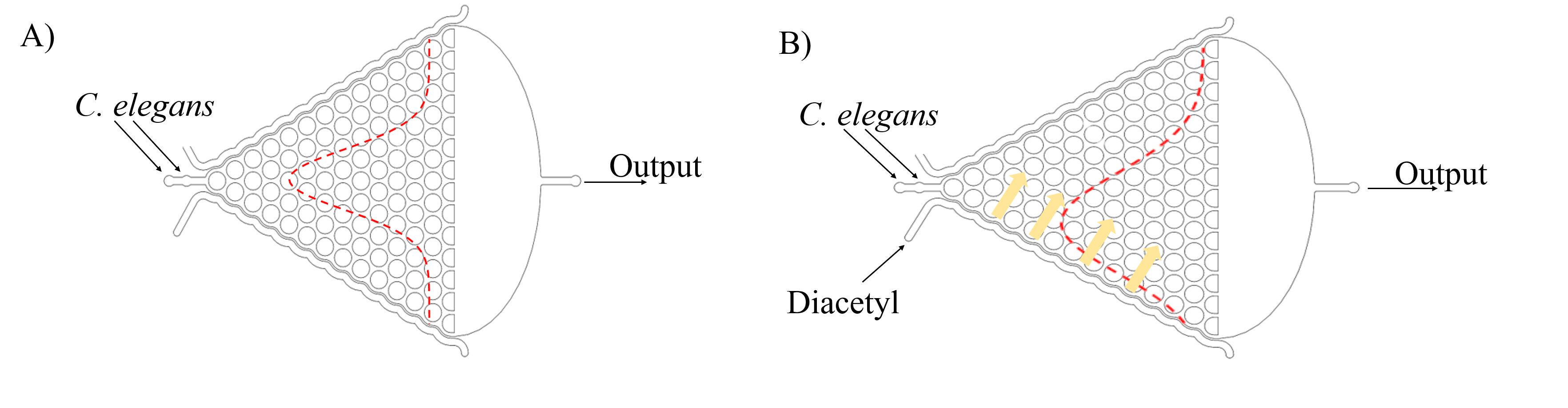
Then, we can assume that C. elegans is just like balls in the Galton board. The force of slow buffer flow acting on worms is the same as the gravity acting on balls. Both of the distribution is Gaussian distribution. Given that we need to make sure that injecting the target genes in C. elegans will not affect its olfactory receptor neuron pair, we injected diacetyl (2-nonanone) that C. elegans prefers (repulse) into the right (left) channel to make a concentration gradient on Gaussian chip. Because of the gradient, worms tend to move to the side filled with diacetyl, causing Gaussian distribution changed. (Fig. 6)
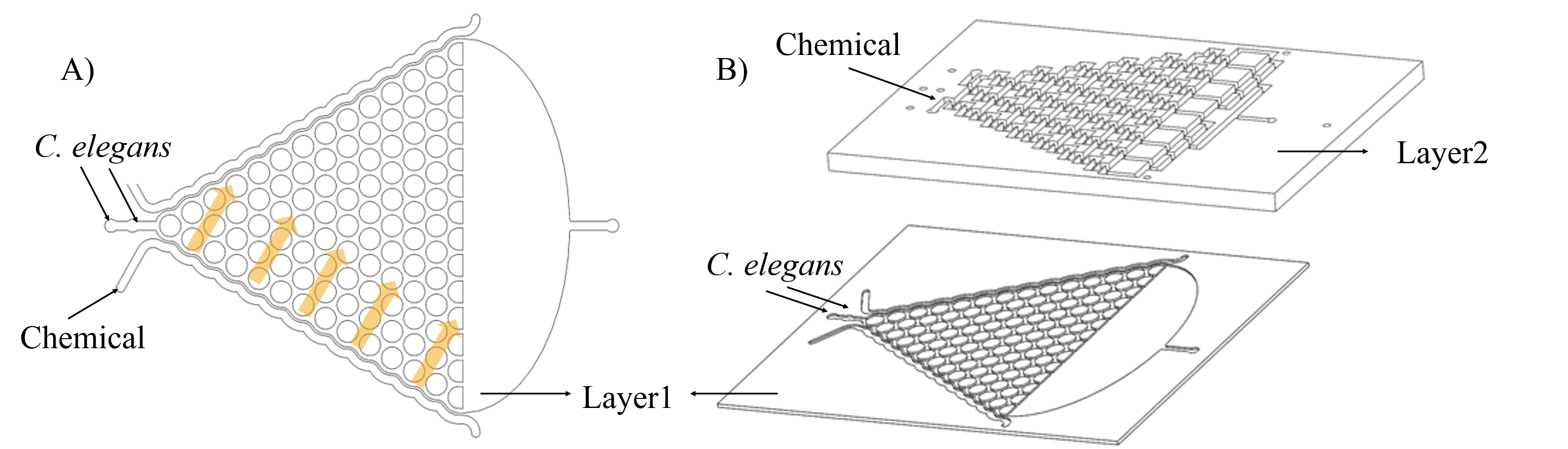
In order to make a concentration gradient, we come up with two methods to get it. (Fig. 7)
In order to simulate the process of diffusion, we make a diffusion model to guide us. More details. See Details.
Both of those methods could be carried out theoretically. But in the process of experiment, we find the method 1 (Inject chemicals into the side of layer 1) is better.
3 The Immobilization Chip
After studying worms’ group behaviors and proving their olfactory neurons are not being affected by exogenous gene, we could study their individual neuron activity and behavioral response. Traditionally, anesthetics and glues are utilized to immobilize worms. However worms will be damaged in these and it will create difficulties in studying its behavioral response. Thus, we designed two kinds of microfluidic chips to allow high-resolution microscopic imaging on chip without damaging for worms. (Fig. 8)
The first one is used to research the imaging of neuronal response, called worm clamps or traps[3].Worms are trapped in the wedge-shaped channel. Trapped worms can then be released by reversing the flow. Traps allow worms’ movement in x-y plane but restrict it in z direction, and it is also utilized to study the contraction and elongation of their heads and research the neuronal activity by detecting calcium indicator GEM-GECO in imaging software.
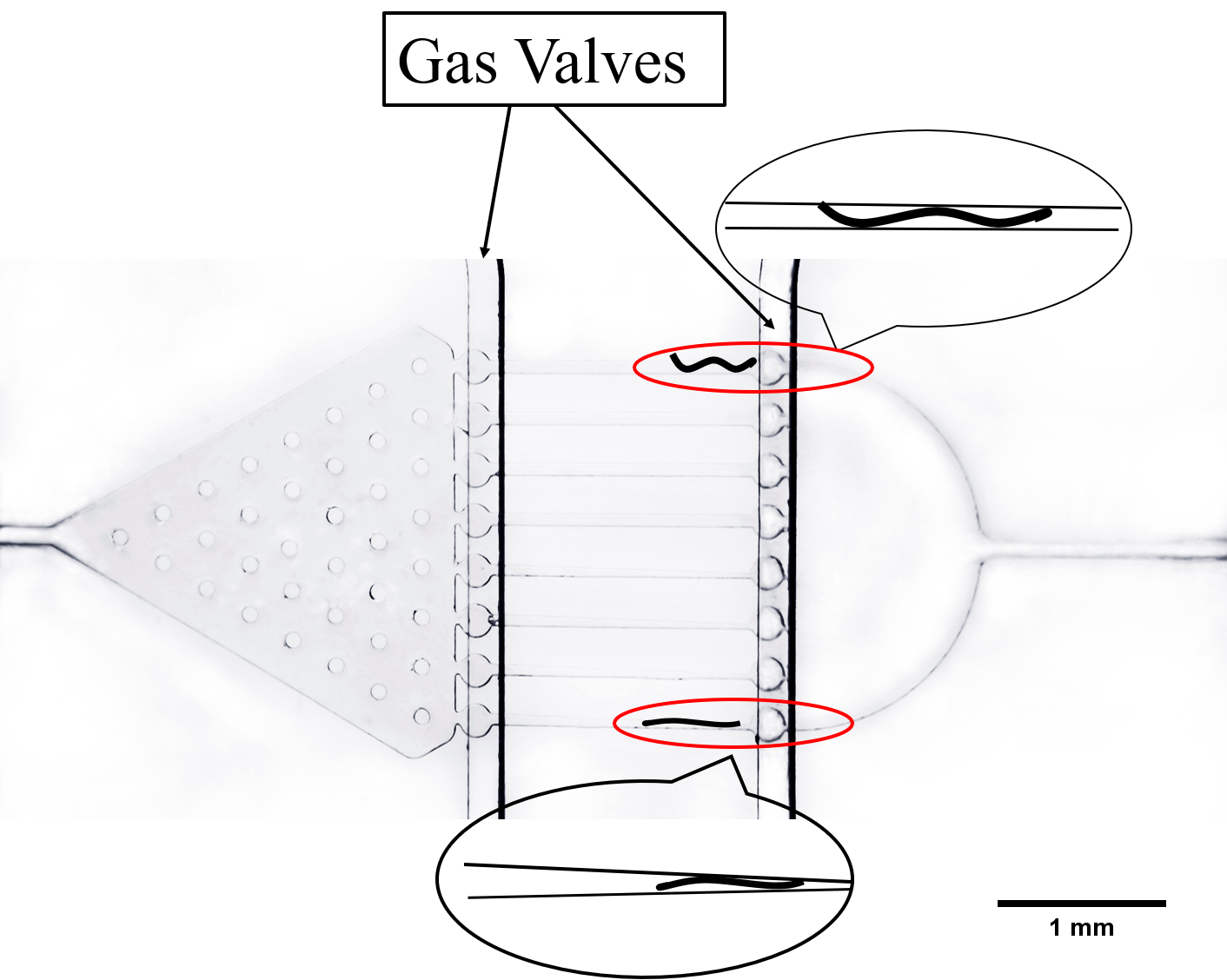
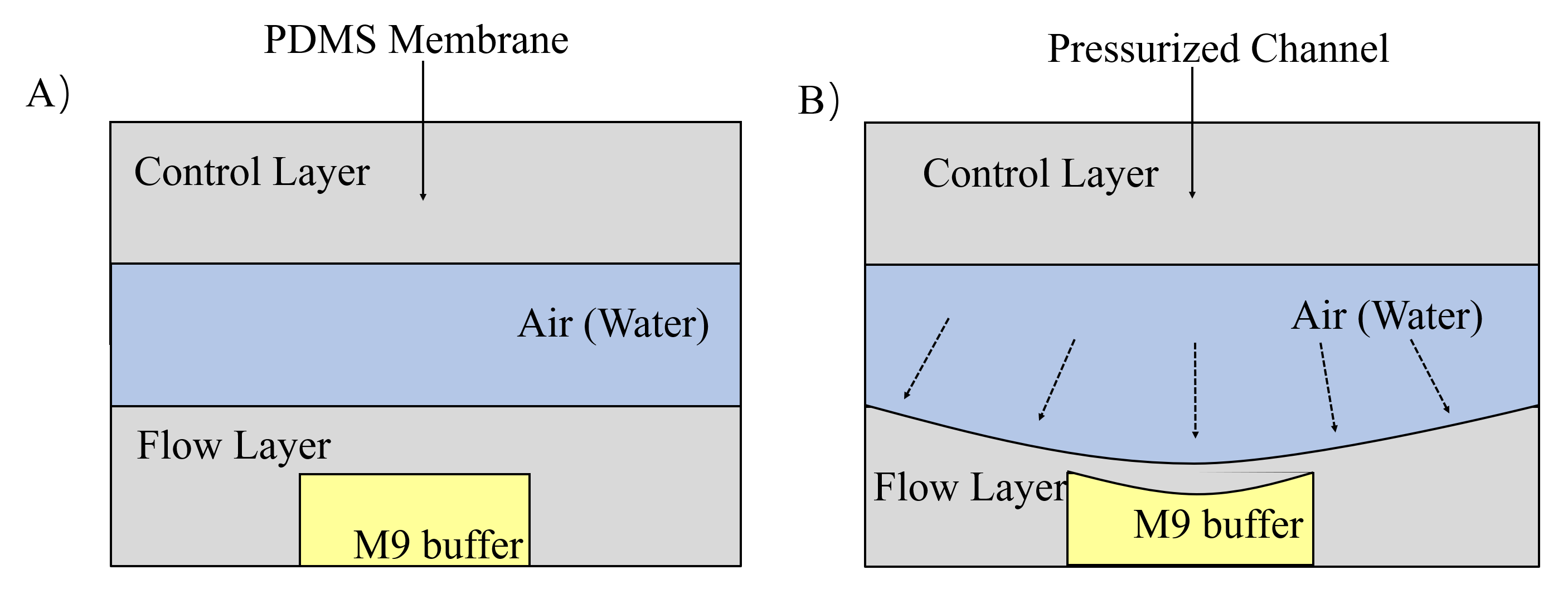
The next channel is compressed and rectangular, called parallel channel. In order to immobilize worms in this channel, we design two gas valves to block the exit and entry by compressing PDMS (a flexible and Easily deformed material) under the pressure made by water or air. (Fig. 9) [4]
The contents above are our all hardware parts in microfluidics. You can learn more details in results. (https://2017.igem.org/Team:SUSTech_Shenzhen/Results)
References
- ↑ Albrecht, D.R., and Bargmann, C.I. (2011). High-content behavioral analysis of Caenorhabditis elegans in precise spatiotemporal chemical environments. Nat. Methods 8, 599-605.
- ↑ Bean machine. (2017, October 5). In Wikipedia, The Free Encyclopedia. Retrieved 12:46, October 22, 2017, from https://en.wikipedia.org/w/index.php?title=Bean_machine&oldid=803992086
- ↑ Hulme, S. E., Shevkoplyas, S. S., Apfeld, J., Fontana, W., & Whitesides, G. M. (2007). A microfabricated array of clamps for immobilizing and imaging C. elegans. Lab on A Chip, 7(11), 1515.
- ↑ Unger, M.A., Chou, H.P., Thorsen, T., Scherer, A., and Quake, S.R. (2000). Monolithic microfabricated valves and pumps by multilayer soft lithography. Science 288, 113-116.