Contents
Overview
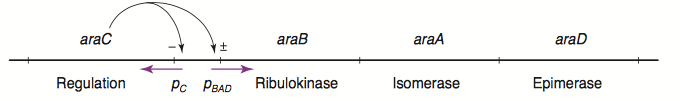
Campylobacter jejuni is the leading cause of food poisoning in the UK, and is commonly found on uncooked poultry. Our team aims to develop a functional biosensor to detect Campylobacter jejuni on contaminated surfaces. We aim to engineer E.coli to produce GFP in response to the presence of xylulose – a sugar found in the outer capsule of C. jejuni. This report focuses on utilization of the L-arabinose sensing AraC regulatory protein and its regulated pBAD promoter to construct one of the parts of the final detection system. The regulatory protein AraC binds to operator sequences in the pBAD promoter and activates transcription of downstream genes only when bound to its substrate sugar (L-arabinose). Here we performed saturation mutagenesis on the araC gene to construct AraC mutant library with four randomized ligand-binding pocket amino acid residues corresponding to positions 8, 24, 80 and 82. The AraC mutant library was subjected to fluorescence-based screening using a GFP reporter. Due to unavailability of xylulose we tested for mutants responsive to xylose, decanal or no effector. We isolated and sequenced 4 mutants which showed altered effector specificity. Our results demonstrate an approach that allows to engineer a regulatory protein to respond to an effector molecule of interest and represent steps towards developing a functional biosensor to detect C. jejuni on contaminated surfaces.
Background
Campylobacter jejuni is a bacterial pathogen found in animal gastrointestinal tract, and is mostly associated with chickens. It is responsible for a vast number of food poisoning cases around the world. Current detection systems for the pathogen are time-consuming, expensive and inaccessible for everyday users. Our team aims to develop a functional biosensor by engineering Escherichia coli to detect xylulose - a sugar naturally present in the outer capsule of Campylobacter jejuni [1]. The sugar can be easily released from the capsule by contact with acid. Our team aims to construct two alternative regulatory systems as potential components of the biosensor. One of them exploits the MtlR operon, naturally found in Pseudomonas fluorescens. MtlR protein activates transcription of downstream genes upon binding to xylulose. Parts of the operon can be transformed into E. coli and utilized in such a way that in contact with xylulose, MtlR protein activates transcription of a gene producing a detectable signal, such as fluorescence from GFP. However, this approach may have some disadvantages. Coming from a different organism, the MtlR protein may not function in E. coli cells as expected. Furthermore, the MtlR operon has not been previously exploited in biosensors and may not show sufficient specificity to xylulose. Hence, we aim to construct an alternative regulatory system which exploits the L-arabinose operon.
Aims
- Aim 1
- Sub-aim 1
- Sub-aim 2
- Aim 2
- Sub-aim 1
- Sub-aim 2
Materials and Methods
Condition set up
Sample preparation
- 1
- 2
- 3